
以三(4-咪唑基苯基)胺为配体的镉配合物的合成、晶体结构和荧光性质
English
Syntheses, Crystal Structures and Photoluminescent Properties of Two Cadmium(II) Coordination Polymers Constructed from Tris(4-imidazolylphenyl)amine
-
As an emerging multifunctional solid crystalline materials over the last two decades, metal-organic frameworks (MOFs) have been a hotspot not only for the diversity of architectures and fascinating topol-ogies but also for potential applications in gas storage and separation, heterogeneous catalysis, fluorescence, chemical sensing, magnetism, electrical energy storage, and so forth[1-7]. Although much progress has been achieved in this field over the past years, it is still a challenge for us to fabricate the desired MOFs with expected structures and properties, because there are varied factors that can affect the structure and property of MOFs[8-12]. From the previously reported studies, it has been demonstrated that organic ligands are crucial in determining the formation of definite MOFs[13-15]. Among various organic ligands, the N-donor ligands as good candidates for the construction of coordination polymers, have aroused a good deal of interests from chemists because of their diversities in coordination modes and conformations[16-17]. It should be noted that, to date, the imidazole-containing N-donor ligands such as 1, 4-di(1H-imidazol-1-yl)benzene, 4, 4′-di(1H-imid-azol-1-yl)biphenyl, 1, 3, 5-tri(1H-imidazol-1-yl)benzene, 3, 3′, 5, 5′-tetra(1H-imidazol-1-yl)biphenyl and 1, 3-di(1H-imidazol-1-yl)benzene, have been widely used in the construction of various coordination polymers[18-22]. However, the coordination polymers constructed by tri(imidazole) ligands are still in its infancy, although some intriguing examples have been reported[23-27]. Tris(4-imidazolylphenyl)amine (TIPA), as a tridentate brid-ging ligand, is rarely used in the construction of coor-dination networks. The TIPA ligand possesses three Ph-imidazole (Ph=phenyl) arms with conformational and geometrical flexibility. The Ph-imidazole arms can rotate freely and adjust itself sterically around the central N moiety when coordinating to the metals[28-32].
In recent years, a number of coordination polymers with various structural types and topological features have been documented[33-35]. In this regard, entangled architectures, including interweaving, polyknotting, polythreading, interdigitation and polycatenation, have been deliberately designed and extensively discussed in several comprehensive reviews[36-37]. Generally, the topological architectures of the coordination polymers can be controlled by the deliberate design and judicious choice of the organic ligands and coordination geometries of the metals[38-39]. In this aspect, the stru-ctural features of the organic ligands, such as shape, functionality, flexibility, symmetry, length, and substituent group, can influence structure types of the coordination polymers directly[40-44]. In this work, two coordination p olymers, [CdI(TIPA)(CDC)0.5]n (1) and {[Cd(TIPA)(MPDA)]·H2O}n (2), have been synthesized by using TIPA ligand and different carboxylate anions, where H2CDC=1, 4-cyclohexanedicarboxylic acid and H2MPDA=5-methylisophthalic acid. The effects of anions on their complex structures are unraveled in detail. Further, the photoluminescent properties of two complexes have also been studied.
1. Experimental
1.1 Materials and general methods
All chemicals and solvents were of reagent grade and used as received without further purification. The TIPA ligand was synthesized according to the reported method[30]. Elemental analyses (C, H and N) were performed on a Vario EL Ⅲ elemental analyzer. Infrared spectra were recorded on KBr discs using a Nicolet Avatar 360 spectrophotometer in the range of 4000~400 cm-1. Thermogravimetric analyses (TGA) were performed on a Netzsch STA-409PC instrument in flowing N2 with a heating rate of 10 ℃·min-1. The luminescent spectra for the powdered solid samples were measured at room temperature on a Horiba FluoroMax-4P-TCSPC fluorescence spectrophotometer with a xenon arc lamp as the light source. In the measurements of emission and excitation spectra the pass width is 5 nm. All the measurements were carried out under the same experimental conditions.
1.2 Synthesis of [CdI(TIPA)(CDC)0.5]n (1)
A mixture containing CdI2 (36.6 mg, 0.1 mmol), H2CDC (17.2 mg, 0.1 mmol), and TIPA (44.3 mg, 0.1 mmol) in DMF/H2O (1:1, V/V) solvent (10 mL) was sealed in a Teflon-lined stainless steel container and heated at 120 ℃ for 3 days. After being cooled down to room temperature, colorless block crystals of 1 were obtained in 57% yield based on TIPA. Anal. Calcd. for C31H25IN7O2Cd(%): C, 48.55; H, 3.29; N, 12.79. Found(%): C, 48.47; H, 3.31; N, 12.77. IR (KBr, cm-1): 3 601 (m), 3 089 (w), 1 581 (s), 1 519 (s), 1 364 (s), 1 307 (m), 1087 (m), 1 015 (w), 966 (w), 811 (m), 746 (m), 697 (w), 657 (w), 543 (w).
1.3 Synthesis of {[Cd(TIPA)(MPDA)]·H2O}n (2)
Complex 2 was prepared by a process similar to that yielding complex 1 at 120 ℃ by using CdI2 (36.6 mg, 0.1 mmol), H2MPDA (17.2 mg, 0.1 mmol), and TIPA (44.3 mg, 0.1 mmol) in DMF/H2O (1:1, V/V) solvent (10 mL). Colorless block crystals of 2 were collected by filtration and washed with water and ethanol several times with a yield of 53% based on TIPA ligand. Anal. Calcd. for C36H29N7O5Cd(%): C, 57.49; H, 3.89; N, 13.04. Found(%): C, 57.44; H, 3.91; N, 13.01. IR (KBr, cm-1): 3 416 (m), 3 091 (m), 1 623 (m), 1 553 (m), 1519 (s), 1 431 (m), 1 381 (s), 1 271 (m), 1 032 (m), 913 (m), 828 (w), 734 (s), 654 (m), 542 (w).
1.4 X-ray crystallography
Two block single crystals with dimensions of 0.27 mm×0.25 mm×0.22 mm for 1 and 0.25 mm×0.22 mm×0.19 mm for 2 were mounted on glass fibers for measure-ment, respectively. X-ray diffraction intensity data were collected on a Bruker APEX Ⅱ CCD diffractometer equipped with a graphite-monochro-matic Mo Kα radiation (λ=0.071 073 nm) using the φ-ω scan mode at 293(2) K. The diffraction data were integrated by using the SAINT program[45], which was also used for the intensity corrections for the Lorentz and polarization effects. Semi-empirical absorption corrections were applied using the SADABS program[46]. The structures were solved by direct methods using SHELXS-2014[47] and all the non-hydrogen atoms were refined anisotropically on F2 by the full-matrix least-squares technique with the SHELXL-2014[48] crystallo-graphic software package. The hydrogen atoms, except those of water molecules, were generated geometrically and refined isotropically using the riding model. The pertinent crystallographic data collection and structure refinement parameters are presented in Table 1 and selected bond lengths and angles are listed in Table 2.
表 1
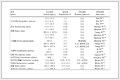 表 1 Crystal data and structure refinement for 1 and 2Table 1. Crystal data and structure refinement for 1 and 2
表 1 Crystal data and structure refinement for 1 and 2Table 1. Crystal data and structure refinement for 1 and 2Complex 1 2 Formula C31H25N7O21Cd C36H29N7O5Cd Formula weight 766.88 752.06 Temperature 293(2) 293(2) Crystal system Monoclinic Monoclinic Space group C2/c P21/c a / nm 2.355 9(2) 1.019 30(9) b/ nm 1.447 25(13) 1.840 95(17) c / nm 1.866 88(17) 1.885 08(17) β/(°) 94.745(2) 104.450(2) V/ nm3 6.343 5(10) 3.425 4(5) Z 8 4 Dc/(g·cm-3) 1.606 1.458 Absorption coefficient/mm 1.701 0.690 θ range / (°) 2.90~25.10 2.86~25.50 F(000) 3 016 1 528 Reflection collected 36 605 50 446 Independent reflection 5 655 (RInt=0.050 5) 6 367 (Rint=0.051 9) Reflection observed [I > 2σ(I)] 4 821 5 564 Data, restraint, parameter 5 655, 0, 398 6 367, 55, 480 Goodness-of-fit on F2 1.043 1.124 R1, wR2 [I > 2σ(I)] 0.033 1, 0.082 7 0.058 8, 0.123 1 R1, wR2(all data) 0.041 7, 0.090 0 0.068 7, 0.128 5 (Δρ)max, (Δρ)min/ (e·nm-3) 1 117, -933 1 042, -895 表 2
 表 2 Selected bond lengths(nm) and angles(°) for 1 and 2Table 2. Selected bond lengths(nm) and angles(°) for 1 and 2
表 2 Selected bond lengths(nm) and angles(°) for 1 and 2Table 2. Selected bond lengths(nm) and angles(°) for 1 and 21 Cd(1)-O(1) 0.223 O(4) Cd(1)-N(4)#2 0.236 9(4) Cd(1)-N(2) 0.229 6(3) Cd(1)-I(1) 0.279 34(4) Cd(1)-N(6)#1 0.235 3(3) O(1)-Cd(1)-N(2) 139.34(15) N(6)#1-Cd(1)-N(4)#2 168.09(14) O(1)-Cd(1)-N(6)#1 88.76(16) O(1)-Cd(1)-I(1) 106.51(12) N(2)-Cd(1)-N(6)#1 88.79(13) N(2)-Cd(1)-I(1) 114.14(9) O(1)-Cd(1)-N(4)#2 84.19(17) N(6)#1-Cd(1)-I(1) 94.21(9) N(2)-Cd(1)-N(4)#2 90.31(14) N(4)#2-Cd(1)-I(1) 96.99(10) 2 Cd(1)-O(1) 0.227 6(4) Cd(1)-O(4)#5 0.237 4(4) Cd(1)-N(3) 0.229 4(5) Cd(1)-N(5)#6 0.238 3(5) Cd(1)-N(7)#4 0.229 7(5) Cd(1)-O(3)#5 0.253 1(4) O(1)-Cd(1)-N(3) 95.18(18) N(7)#4-Cd(1)-N(5)#6 89.2(2) O(1)-Cd(1)-N(7)#4 136.02(16) O(4)#5-Cd(1)-N(5)#6 85.44(19) N(3)-Cd(1)-N(7)#4 92.35(18) O(1)-Cd(1)-O(3)#5 136.2O(16) O(1)-Cd(1)-O(4)#5 84.15(15) N(3)-Cd(1)-O(3)#5 90.98(18) N(3)-Cd(1)-O(4)#5 89.94(18) N(7)#4-Cd(1)-O(3)#5 86.76(16) N(7)#4-Cd(1)-O(4)#5 139.21(16) O(4)#5-Cd(1)-O(3)#5 52.47(15) O(1)-Cd(1)-N(5)#6 87.3(2) N(5)#6-Cd(1)-O(3)#5 83.9(2) N(3)-Cd(1)-N(5)#6 174.5(2) Symmetry codes: #1: -x+1, -y+2, -z+1; #2: x, -y+1, z-1/2 for 1; #4: -x+1, y+1/2, -z+1/2; #5: x+1, y, z; #6: x, y+1, z for 2. CCDC: 1557335, 1; 1557336, 2.
2. Results and discussion
2.1 Crystal structure
Single crystal X-ray diffraction analysis revealed that complex 1 crystallizes in the monoclinic space group C2/c. The asymmetric unit of complex 1 contains one crystallography independent Cd(Ⅱ) ion, one unique TIPA ligand, one iodide ion and a half CDC2- anion which locates at an inversion center as illustrated in Fig. 1. The Cd(Ⅱ) ion is five-coordinated by three nitrogen atoms from three TIPA ligands, one iodide ion and one oxygen atom in a distorted square pyramidal geometry with a τ value of 0.479[49]. The Cd-N bond lengths vary from 0.229 6(3) to 0.236 9(4) nm, and N-Cd-N angles range from 88.79(13)° to 168.09(14)°.
图 1
Each TIPA ligand bridges three Cd(Ⅱ) ions to generate a highly undulant 2D sheet with a thickness of ca. 1.13 nm (Fig. 2). From a topological viewpoint, the sheet can be considered as a 3-connected network with a Schlfli symbol of (4.82). There are two kinds of large windows of Cd2(TIPA)2 and Cd4(TIPA)4 in the resulting layers. The Cd2(TIPA)2 window is built up by two Cd(Ⅱ) ions and four Ph-imidazole arms of two different TIPA ligands with the dimension of 1.202 nm×1.529 nm, while the Cd4(TIPA)4 unit is built up by four Cd(Ⅱ) ions and eight Ph-imidazole arms of four ligands with the dimension of 1.126 nm×3.335 nm. It is noteworthy that each Cd4(TIPA)4 unit is divided into two Cd2(TIPA)2(CDC) subunit by CDC anion linking two Cd(Ⅱ) ions to give an undulate 2D layer. Topolo-gically, the TIPA ligand can be considered as a 3-connected node, the CDC anions can be considered as linkers, and Cd(Ⅱ) ions can be regarded as 4-connected nodes. Therefore, the 2D layer of 1 is a (3, 4)-connected topology with Schläfli symbol of (4.52)(4.53.72) (Fig. 3).
图 2
图 3
The most fascinating structural feature of 1 is that two such layers are interlaced each other in a parallel fashion to give a 2D→2D polyrotaxane network (Fig. 4). The Cd2TIPA2 windows of each layer are passed by CDC anions of adjacent layer, and each Cd2(TIPA)2(CDC) unit of each layer is threaded through by one armed rod of the TIPA ligand from adjacent layers. More interestingly, the neighboring 2D polyrotaxane sheets are parallel with each other.
图 4
When the ligand H2MPDA was used instead of H2CDC to react with Cd(Ⅱ) salts under hydrothermal conditions, Complex 2 was isolated. Complex 2 crysta-llizes in the monoclinic space group P21/c. The asym-metric unit consists of one Cd(Ⅱ) ion, one TIPA ligand, and one lattice water molecule. The local coordination geometry around the Cd(Ⅱ) ion is depicted in Fig. 5. The Cd(Ⅱ) ion adopts a distorted octahedral coordina-tion sphere that is defined by three oxygen atoms from two distinct MPDA2- anions and three nitrogen atoms from three different TIPA ligands; thus, the Cd(Ⅱ) ions can be considered as 5-connecting nodes. Each TIPA links three Cd(Ⅱ) ions, acting as a 3-connecting node to form a 1D ladder-like chain. Two carboxylate groups of the MPDA2- anions adopting monodentate and bide-ntate chelate coordination modes bridge two Cd(Ⅱ) ions, and MPDA2- joins adjacent parallel chains as a pillar connector to form a 2D grid like (4, 4) bilayer (Fig. 6). The Schläfli symbol for this binodal net is (42·67·8)(42·6). Viewed from the b axis, there exist rectangular channels with a cross section of approxi-mately 2.04 nm×1.09 nm (excluding van der Waals radii). The channel is so large that the two other equivalent bilayers can be accommodated in that channel. This may be the mainly reason to form the 2D→3D parallel entangled structure. Upon interpene-tration, Complex 2 just contains a small solvent accessible void space of 2.8% of the total crystal volume, according to a calculation performed using PLATON[50].
图 5
图 6
The topological feature of Complex 2 is most unusual because it is a rarely observed bilayer motif which is parallel-parallel catenated with two other equivalent adjacent ones to form a 3D supramolecular structure (Fig. 7). To our knowledge, this (3, 5)-connected bilayer is yet to be reported. Compared to 2D→3D polycatenation systems in parallel-parallel inclined fashion or parallel-parallel highly undulating fashion, fewer examples of 2D→3D parallel entangled structures have been observed[51-52].
图 7
2.2 FT-IR spectra
The IR spectra of 1 and 2 show the absence of the characteristic bands at around 1 700 cm-1 attri-buted to the protonated carboxylate group, which indicates that the complete deprotonation of H2CDC and H2MPDA ligands upon reaction with Cd(Ⅱ) ion. The presence of vibrational bands of 1 650~1 550 cm-1 are characteristic of the asymmetric stretching of the deprotonated carboxylic groups of CDC2- and MPDA2- anions. The difference between asymmetric and symmetric carbonyl stretching frequencies (Δν=νasym-νsym) was used to fetch information on the metal-carboxylate binding modes. complex 1 showed a pairs of νasym and νsym frequencies at 1 584, 1 354 cm-1 corresponding to the carbonyl functionality of dicar-boxylate ligand indicating a symmetric monodentate coordination mode(Δν=230 cm-1). Complex 2 shows two pairs of νasym and νsym frequencies at 1 623, 1 431 cm-1(Δν=192 cm-1) and 1 553, 1 381 cm-1 (Δν=172 cm-1) for the carbonyl functionality indicating two coordination modes as observed in the crystal structure. OH stretching broad bands at 3 416 cm-1 for 2 are attributable to the lattice water. The bands in the region of 640~1 250 cm-1 are attributed to the -CH-in-plane or out-of-plane bend, ring breathing, and ring deformation absorptions of benzene ring, respectively. The IR spectra exhibit the characteristic peaks of imidazole groups at ca. 1 520 cm-1 [53].
2.3 Thermal stability
The thermal behaviors of complexes 1 and 2 were measured under a dry N2 atmosphere at a heating rate of 10 ℃·min-1 from 25 to 800 ℃ and the TG curves are presented in Fig. 8. The TGA curve of 1 reveals that no obvious weight loss is observed until the temperature reaches 340 ℃. The anhydrous compound decomposes from 340 to 800 ℃, indicating the release of organic components. For 2, the first weight loss of 2.61% (Calcd. 2.39%) occurs in the range of 60~150 ℃, indicating the loss of one free water molecules. Then, the framework of 2 decomposes gradually above 270 ℃.
图 8
2.4 Photoluminescent properties
Photoluminescence properties of Cd(Ⅱ) complexes have attracted intense interest due to their potential applications in photochemistry, chemical sensors, and electroluminescent display[54-55]. The photoluminescent properties of 1, 2 and TIPA ligand were investigated in solid state at room temperature (Fig. 9). The TIPA ligand exhibits emission band with a maximum at 422 nm upon excitation at 365 nm, which may be assigned to π*→n or π*→π transitions of the ligands[56-57].
图 9
The emission peaks of complexes occur at 429 nm (λex=370 nm) for 1, 482 nm (λex=380 nm) for 2. Under the same experimental conditions, the emission intensities of free H2CDC and H2MPDA are weaker than that of TIPA ligand, so it is considered that it has no significant contribution to the fluorescent emission of the complexes with the presence of TIPA ligand. The emission of 1 can be essentially ascribed to the intraligand fluorescent emission. The red-shift of the emission can be attributed to the ligand coordination to the metal center, which effectively increases the rigidity of the ligand and reduces the loss of energy by radiationless decay. However, the intense blue emission peak at 481 nm (λex=380 nm) for 2 is highly red-shifted with respect to the free TIPA ligands. The intense fluorescence of 2 indicates that a strong interaction exists between the ligand and the Cd(Ⅱ) ion[58-60].
-
-
[1]
Gamage N D H, McDonald K A, Matzger A J. Angew. Chem., Int. Ed., 2016, 55:12099-12103 doi: 10.1002/anie.201606926
-
[2]
Noh T H, Jung O S. Acc. Chem. Res., 2016, 49:1835-1843 doi: 10.1021/acs.accounts.6b00291
-
[3]
Maza W A, Padilla R, Morris A J. J. Am. Chem. Soc., 2015, 137:8161-8168 doi: 10.1021/jacs.5b03071
-
[4]
Manna P, Das S K. Cryst. Growth Des., 2015, 15:1407-1421 doi: 10.1021/cg501787m
-
[5]
He H M, Song Y, Sun F X, et al. Cryst. Growth Des., 2015, 15:2033-2038 doi: 10.1021/acs.cgd.5b00229
-
[6]
Li S L, Xu Q. Energy Environ. Sci., 2013, 6:1656-1683 doi: 10.1039/c3ee40507a
-
[7]
Zheng J, Wu M Y, Jiang F L, et al. Chem. Sci., 2015, 6:3466-3470 doi: 10.1039/C5SC00213C
-
[8]
Long L S. CrystEngComm, 2010, 12:1354-1365 doi: 10.1039/b921146b
-
[9]
Tan Y X, He Y P, Zhang Y, et al. CrystEngComm, 2013, 15: 6009-6014 doi: 10.1039/c3ce40677f
-
[10]
Huang S Y, Li J Y, Li J Q, et al. Dalton Trans., 2014, 43: 5260-5264 doi: 10.1039/c3dt53123f
-
[11]
Zhang J W, Kan X M, Liu B Q, et al. Chem. Eur. J., 2015, 21:16219-16228 doi: 10.1002/chem.201502203
-
[12]
Zhao X L, Sun W Y. CrystEngComm, 2014, 16:3247-3258 doi: 10.1039/c3ce41791c
-
[13]
吴琪, 苏志, 王红艳.无机化学学报, 2017, 33(10): 1889-1895 doi: 10.11862/CJIC.2017.196WU Qi, SU Zhi, WANG Hong-Yan, et al. Chinese J. Inorg. Chem., 2017, 33(10): 1889-1895 doi: 10.11862/CJIC.2017.196
-
[14]
Han L J, Yan W, Chen S G, et al. Inorg. Chem., 2017, 56: 2936-2940 doi: 10.1021/acs.inorgchem.6b03075
-
[15]
He Y F, Chen D M, Xu H, et al. CrystEngComm, 2015, 17: 2471-2478 doi: 10.1039/C4CE02380C
-
[16]
Huang Y Q, Chen H Y, Li Z G, et al. Inorg. Chim. Acta, 2017, 466:71-77 doi: 10.1016/j.ica.2017.05.039
-
[17]
Dong X Y, Si C D, Fan Y, et al. Cryst. Growth Des., 2016, 16:2062-2073 doi: 10.1021/acs.cgd.5b01734
-
[18]
Ke C H, Lin G R, Kuo B C, et al. Cryst. Growth Des., 2012, 12:3758-3765 doi: 10.1021/cg300559p
-
[19]
Wang F, Ke X H, Zhao J B, et al. Dalton Trans., 2011, 40: 11856-11865 doi: 10.1039/c1dt11130b
-
[20]
Su Z, Fan J, Okamura T A, et al. Cryst. Growth Des., 2010, 10:1911-1922 doi: 10.1021/cg100020t
-
[21]
Luo L, Wang P, Xu G C, et al. Cryst. Growth Des., 2012, 12: 2634-2645 doi: 10.1021/cg300220q
-
[22]
Schlechte L, Bon V, Grunker R, et al. Polyhedron, 2012, 44: 179-186 doi: 10.1016/j.poly.2012.06.065
-
[23]
Su Z, Xu J, Fan J, et al. Cryst. Growth Des., 2009, 9:2801-2811 doi: 10.1021/cg900059m
-
[24]
Fan J, Sun W Y, Okamura T A, et al. Inorg. Chem., 2003, 42:3168-3175 doi: 10.1021/ic0206847
-
[25]
Fan J, Gan L, Kawaguchi H, et al. Chem. Eur. J., 2003, 9: 3965-3973 doi: 10.1002/(ISSN)1521-3765
-
[26]
Liu H K, Sun W Y, Ma D J, et al. Chem. Commun., 2000: 591-592
-
[27]
Liu F H, Chen W Z, You X Z. J. Solid State Chem., 2002, 169:199-207 doi: 10.1016/S0022-4596(02)00063-4
-
[28]
Wu H, Liu H Y, Liu Y Y, et al. Chem. Commun., 2011, 47: 1818-1820 doi: 10.1039/C0CC04724D
-
[29]
Yao X Q, Cao D P, Hu J S, et al. Cryst. Growth Des., 2011, 11:231-239 doi: 10.1021/cg1011764
-
[30]
Wu H, Liu H Y, Liu B, et al. CrystEngComm, 2011, 13: 3402-3407 doi: 10.1039/c0ce00908c
-
[31]
Wu H, Liu H Y, Yang J, et al. Cryst. Growth Des., 2011, 11: 2317-2324 doi: 10.1021/cg200005q
-
[32]
Wu H, Liu B, Yang J, et al. CrystEngComm, 2011, 13:3661-3664 doi: 10.1039/c1ce05379e
-
[33]
Alezi D, Spanopoulos I, Tsangarakis C, et al. J. Am. Chem. Soc., 2016, 138:12767-12770 doi: 10.1021/jacs.6b08176
-
[34]
Gu J Z, Liang X X, Cai Y, et al. Dalton Trans., 2017, 46: 10908-10925 doi: 10.1039/C7DT01742A
-
[35]
Duan J G, Higuchi M, Kitagawa S. Inorg. Chem., 2015, 54: 1645-1649 doi: 10.1021/ic502643m
-
[36]
Batten S R. CrystEngComm, 2001, 3:67-72 doi: 10.1039/b102400k
-
[37]
Carlucci L, Ciani G, Proserpio D M. Coord. Chem. Rev., 2003, 246:247-289 doi: 10.1016/S0010-8545(03)00126-7
-
[38]
Dinca M, Yu A F, Long J R. J. Am. Chem. Soc., 2006, 128: 8904-8913 doi: 10.1021/ja061716i
-
[39]
Zhang L P, Ma J F, Yang J, et al. Inorg. Chem., 2010, 49: 1535-1550 doi: 10.1021/ic9019553
-
[40]
Reger D L, Wright T D, Semeniuc R F, et al. Inorg. Chem., 2001, 40:6212-6219 doi: 10.1021/ic010574k
-
[41]
Zaman M B, Smith M D, zur Loye H C. Chem. Commun., 2001:2256-2257 https://www.researchgate.net/profile/Md_Zaman3/publication/11151006_Two_different_one-dimensional_structural_motifs_in_the_same_coordination_polymer_A_novel_interpenetration_of_infinite_ladders_by_bundles_of_infinite_chains/links/5526c23c0cf2e486ae40c2d9.pdf
-
[42]
Kitaura R, Seki K, Akiyama G, et al. Angew. Chem., Int. Ed., 2003, 42:428-431 doi: 10.1002/anie.200390130
-
[43]
Kitaura R, Fujimoto K, Noro S, et al. Angew. Chem., Int. Ed., 2002, 41:133-135 doi: 10.1002/1521-3773(20020104)41:1<>1.0.CO;2-5
-
[44]
Qi Y, Luo F, Che Y X, et al. Cryst. Growth Des., 2008, 8: 606-611 doi: 10.1021/cg700758c
-
[45]
SMART and SAINT, Program for Data Extraction and Reduction, Bruker AXS, Inc., Madison, Wisconsin, USA, 2002.
-
[46]
Sheldrick G M. SADABS, Program for Empirical Absorption Correction of Area Detector Data, University of Göttingen, Germany, 2003.
-
[47]
Sheldrick G M. SHELXS-2014, Program for Crystal Structure Solution, University of Göttingen, Germany, 2014.
-
[48]
Sheldrick G M. SHELXL-2014, Program for the Refinement of Crystal Structure, University of Göttingen, Germany, 2014.
-
[49]
Addison A W, Rao T N. J. Chem. Soc. Dalton Trans., 1984: 1349-1356
-
[50]
Spek A L. J. Appl. Crystallogr., 2003, 36:7-13 doi: 10.1107/S0021889802022112
-
[51]
Zhang J, Chew E, Chen S, et al. Inorg. Chem., 2008, 47: 3495-3497 doi: 10.1021/ic800195u
-
[52]
Wu H, Ma J F, Liu Y Y, et al. CrystEngComm, 2011, 13: 7121-7128 doi: 10.1039/c1ce05889d
-
[53]
Nakamoto K. Infrared and Raman Spectra of Inorganic and Coordinated Compounds. 5th Ed. New York: John Wiley & Sons, 1997.
-
[54]
Thirumurugan A, Natarajan S. Dalton Trans., 2004:2923-2928
-
[55]
McGarrah J E, Kim Y J, Hissler M, et al. Inorg. Chem., 2001, 40:4510-4511 doi: 10.1021/ic015559u
-
[56]
Wen L L, Lu Z D, Lin J G, et al. Cryst. Growth Des., 2007, 7:93-99 doi: 10.1021/cg0604982
-
[57]
Lin J G, Zang S Q, Tian Z F, et al. CrystEngComm, 2007, 9: 915-921 doi: 10.1039/b708389k
-
[58]
Liu H J, Tao X T, Yang J X, et al. Cryst. Growth Des., 2008, 8:259-264 doi: 10.1021/cg070276j
-
[59]
Klessinger J M, Michl J. Excited States and Photochemistry of Organic Molecules. New York: VCH, 1995.
-
[60]
徐涵.无机化学学报, 2016, 32(8):1481-1486 doi: 10.11862/CJIC.2016.194XU Han. Chinese J. Inorg. Chem., 2016, 32(8):1481-1486 doi: 10.11862/CJIC.2016.194
-
[1]
-
Table 1. Crystal data and structure refinement for 1 and 2
Complex 1 2 Formula C31H25N7O21Cd C36H29N7O5Cd Formula weight 766.88 752.06 Temperature 293(2) 293(2) Crystal system Monoclinic Monoclinic Space group C2/c P21/c a / nm 2.355 9(2) 1.019 30(9) b/ nm 1.447 25(13) 1.840 95(17) c / nm 1.866 88(17) 1.885 08(17) β/(°) 94.745(2) 104.450(2) V/ nm3 6.343 5(10) 3.425 4(5) Z 8 4 Dc/(g·cm-3) 1.606 1.458 Absorption coefficient/mm 1.701 0.690 θ range / (°) 2.90~25.10 2.86~25.50 F(000) 3 016 1 528 Reflection collected 36 605 50 446 Independent reflection 5 655 (RInt=0.050 5) 6 367 (Rint=0.051 9) Reflection observed [I > 2σ(I)] 4 821 5 564 Data, restraint, parameter 5 655, 0, 398 6 367, 55, 480 Goodness-of-fit on F2 1.043 1.124 R1, wR2 [I > 2σ(I)] 0.033 1, 0.082 7 0.058 8, 0.123 1 R1, wR2(all data) 0.041 7, 0.090 0 0.068 7, 0.128 5 (Δρ)max, (Δρ)min/ (e·nm-3) 1 117, -933 1 042, -895 Table 2. Selected bond lengths(nm) and angles(°) for 1 and 2
1 Cd(1)-O(1) 0.223 O(4) Cd(1)-N(4)#2 0.236 9(4) Cd(1)-N(2) 0.229 6(3) Cd(1)-I(1) 0.279 34(4) Cd(1)-N(6)#1 0.235 3(3) O(1)-Cd(1)-N(2) 139.34(15) N(6)#1-Cd(1)-N(4)#2 168.09(14) O(1)-Cd(1)-N(6)#1 88.76(16) O(1)-Cd(1)-I(1) 106.51(12) N(2)-Cd(1)-N(6)#1 88.79(13) N(2)-Cd(1)-I(1) 114.14(9) O(1)-Cd(1)-N(4)#2 84.19(17) N(6)#1-Cd(1)-I(1) 94.21(9) N(2)-Cd(1)-N(4)#2 90.31(14) N(4)#2-Cd(1)-I(1) 96.99(10) 2 Cd(1)-O(1) 0.227 6(4) Cd(1)-O(4)#5 0.237 4(4) Cd(1)-N(3) 0.229 4(5) Cd(1)-N(5)#6 0.238 3(5) Cd(1)-N(7)#4 0.229 7(5) Cd(1)-O(3)#5 0.253 1(4) O(1)-Cd(1)-N(3) 95.18(18) N(7)#4-Cd(1)-N(5)#6 89.2(2) O(1)-Cd(1)-N(7)#4 136.02(16) O(4)#5-Cd(1)-N(5)#6 85.44(19) N(3)-Cd(1)-N(7)#4 92.35(18) O(1)-Cd(1)-O(3)#5 136.2O(16) O(1)-Cd(1)-O(4)#5 84.15(15) N(3)-Cd(1)-O(3)#5 90.98(18) N(3)-Cd(1)-O(4)#5 89.94(18) N(7)#4-Cd(1)-O(3)#5 86.76(16) N(7)#4-Cd(1)-O(4)#5 139.21(16) O(4)#5-Cd(1)-O(3)#5 52.47(15) O(1)-Cd(1)-N(5)#6 87.3(2) N(5)#6-Cd(1)-O(3)#5 83.9(2) N(3)-Cd(1)-N(5)#6 174.5(2) Symmetry codes: #1: -x+1, -y+2, -z+1; #2: x, -y+1, z-1/2 for 1; #4: -x+1, y+1/2, -z+1/2; #5: x+1, y, z; #6: x, y+1, z for 2. -

 扫一扫看文章
扫一扫看文章
计量
- PDF下载量: 5
- 文章访问数: 1958
- HTML全文浏览量: 125

 下载:
下载:









 下载:
下载:
 下载:
下载:

