
3, 5-二烷基-1, 2, 4-三唑衍生物构筑的两个Cu4I4簇配合物的合成、结构和荧光性质
-
关键词:
- 亚铜碘配合物
- / 1, 2, 4-三唑
- / 晶体结构
- / 荧光性质
English
Syntheses, Crystal Structures and Luminescence Properties of Two Cu4I4 Coordination Polymers Based on 3, 5-Dialkyl-1, 2, 4-triazole
-
Key words:
- copper (I) iodide
- / 1, 2, 4-triazole
- / crystal structures
- / luminescence
-
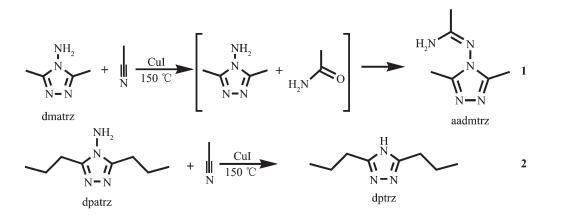
Cuprous halide complexes have been of great interest for their various structure, unique topologies[1-3], as well as attractive properties such as luminescence[4-5], pores[6] and high reactivity in numerous organic and biochemical reactions[7-8]. (CuxXy)x-y (X=halide)[9] have richer aggregates of different geometries: Cu2X2, Cu3X3, Cu3X7, Cu3X8, Cu4X4, Cu4X11, Cu4X9, Cu6X6[10-14], and so on. Neutral cuprous halide clusters show Cu2X2, Cu4X4, and Cu6X6. These aggregates can be linked by organic ligands to form cuprous halide coordination polymers. It has been reported N-donor ligands such as pyrazine, bipyridyl[11], imidazole, triazole[13, 15], tetrazole[9], benzotriazole[16]. To date, a large number of cuprous halide complexes containing triazole ligands have been reported[17-19]. What we are interested in is that cuprous iodide with 4-aminotriazole shows diversities of Cu(Ⅰ) coordination geometry and new 4-aminotriazole ligands are self-assembled by the condensation reaction or the reductive deaminization[20]. Herein, we report the syntheses and crystal structures of two Cu4I4 aggregates complexes {[Cu2(aadmtrz)I2]·CH3CN}n (1) and [Cu2(dptrz)I]n (2), and illustrate that the length of alkyl side chains of 4-aminotriazole has a certain extent influence on the product formations, structures and topologies. The complexes were characterized by X-ray single crystal analysis and studied by various spectroscopic techniques.
1 Experimental
1.1 Materials and methods
All reagents were purchased commercially and used without further purification. Ligands dmatrz and dpatrz were synthesized according to the literature[21]. Elemental analyses (C, H, and N) were performed on an Elementar Vario EL Ⅲ analyzer. FTIR spectra were recorded from KBr pellets in the range of 4 000 ~400 cm-1 on a Bruker Tensor 27 spectrometer. PXRD data were collected in a Bruker D8 Advance X-ray diffractometer using Cu Kα radiation (λ=0.154 059 8 nm) at 30 kV and 15 mA over the 2θ range of 5°~50°. The simulated patterns of 1 and 2 were derived from free Mercury Version 2.2 software. Luminescence spectra were recorded on a CARY Eclipse (Varian, USA) fluorescence spectrophotometer at room temperature.
1.2 Synthesis of complex {[Cu2(aadmtrz)I2]·CH3CN}n (1)
A mixture containing CuI (0.10 g, 0.53 mmol), dmatrz (0.056 g, 0.50 mmol), CH3CN (1 mL) and H2O (5 mL) was sealed in a 23 mL of Teflon-lined stainless steel vessel, which was heated at 150 ℃ for 4 days, and cooled to room temperature. Light-green block crystals of 1 were obtained, and picked out, washed with CH3CN/H2O (1:5, V/V) and dried in air. Yield: 47.5%. Anal. Calcd. for C8H14Cu2I2N6(%): C, 16.71; H, 2.45; N, 14.61. Found(%): C, 16.68; H, 2.51; N, 14.62. IR (cm-1): 3 315 (s), 3 216 (m), 3 004 (w), 2 968 (w), 2 914 (w), 1 622 (s), 1 543 (s), 1 427 (s), 1 373 (m), 1 266 (m), 1 043 (m), 990(m), 891(s), 758(s), 742(m).
1.3 Synthesis of complex [Cu2(dptrz)I]n (2)
The same synthetic procedure as that for 1 was used except that dmatrz was replaced with dpatrz (0.084 g, 0.50 mmol), giving pale-green octahedral crystals 2 in 25.8% yield. Anal. Calcd. for C8H14Cu2 N3I(%): C, 23.65; H, 3.47; N, 10.34, Found(%): C, 23.58; H, 3.44; N, 10.32%. IR (cm-1): 3 318 (s), 3 255(m), 3 198 (s), 2 970 (s), 2 927 (m), 2 880 (m), 2 052 (s), 1 621 (s), 1 544 (s), 1 460 (m), 1 390 (m), 1 343 (w), 1 293 (w), 1 210 (w), 1 072 (w), 946 (m), 889 (w), 715 (m).
1.4 X-ray crystallography
Single-crystal X-ray diffraction data for comp-lexes 1 and 2 were collected in Beijing Synchrotron Radiation Facility (BSRF) beamline 3W1A which mounted with a MARCCD-165 detector (λ=0.072 00 nm) with storage ring working at 2.5 GeV. In the process, the crystals were protected by liquid nitrogen at 100(2) K. Data were collected by the MARCCD and processed using HKL 2000[22]. Absorption correc-tions were applied by using the multi-scan program SCALEPACK[22]. All the structures were solved by the direct methods and refined by the full-matrix least-squares technique using the SHELXL-2014[23] with all non-hydrogen atoms refined anisotropically. Hydrogen atoms attached to C and N atoms were added theoretically and treated as riding on the concerned atoms. The final cycle of full-matrix least-squares refinement was based on observed reflections and variable parameters. Further crystallographic data and structural refinement details are summarized in Table 1. Selected bond lengths and bond angles are given in Table 2.
1 2 Formula C8H14Cu2I2N6 C8H14Cu2N3I Formula weight 575.13 406.20 Crystal system Monoclinic Tetragonal Space group P21/c I41/a a / nm 0.954 7(1) 1.625 3(2) b/ nm 1.187 3(2) 1.625 3(2) c / nm 1.494 9(5) 1.907 8(4) β/(°) 111.23(3) V / nm3 1.579 5(7) 5.039 5(1) Crystal size / mm 0.45×0.40×0.40 0.05×0.04×0.03 Z 4 16 Dc / (g·cm-3) 2.419 2.142 μ/ mm-1 6.59 5.794 F(000) 1 072 3 104 θ range / (°) 2.9~26.0 2.51~25.05 Gocxlness-of-fit on F2 1.41 1.046 Reflection collected, unique 3 075, 3 067 2 234, 1 940 Rint 0.017 0.028 7 R1, wR2[I>2σ(I)] 0.019 1, 0.058 6 0.032 9, 0.089 2 R1, wR2(all data) 0.020 9, 0.072 5 0.038 6, 0.092 8 (Δ/σ)max 0.002 0.004 Largest diff peak and hole / (e • nm-3) 1 180, -1 580 697, -1 880 1 Сu(1)-N(1) 0.201 2(3) Cu(1)-N(4) 0.206 3(3) Cu(1)-I(1) 0.268 9(0) Сu(1)-I(2) 0.267 1(1) Cu(2)-N(2) 0.200 1(3) Cu(2)-I(1) 0.258 0(1) Сu(2)-I(2)ⅰ 0.252 3(1) Сu(1)...Сu(2)ⅰ 0.288 4(1) Сu(2)-Сu(1)ⅰ 0.288 4(1) Cu(2)-I(1)-Cu(1) 81.61(2) Сu(2)ⅰ-1(2)-Сu(1) 67.41(3) N(1)-Cu(1)-N(4) 120.04(1) N(1)-Cu(1)-I(2) 110.45(9) N(4)-Cu(1)-I(2) 103.08(8) N(1)-Cu(1)-I(1) 102.81(9) N(4)-Cu(1)-I(1) 104.16(9) I(2)-Cu(1)-I(1) 116.90(3) N(1)-Cu(1)-Cu(2)ⅰ 88.55(9) N(4)-Cu(1)-Cu(2)ⅰ 149.84(8) I(2)-Сu(1)-Сu(2)ⅰ 53.84(2) I(1)-Сu(1)-Сu(2)ⅰ 75.91(3) N(2)-Cu(2)-I(2)ⅰ 128.18(9) N(2)-Cu(2)-I(1) 105.90(8) I(2) i-Cu(2)-I(1) 122.09(3) N(2)-Cu(2)-Cu(1)ⅰ 121.26(9) I(2)ⅰ-Сu(2)-Сu(1)ⅰ 58.75(2) I(1)-Сu(2)-Сu(1)ⅰ 112.17(2) 2 Сu(1)-N(1) 0.191 5(4) Cu(1)-I(1) 0.249 1(1) Cu(1)…Сu(1)ⅰ 0.254 4(1) Сu(1)-І(1)ⅱ 0.259 9(1) Cu(1)…Cu(1)ⅲ 0.261 1(1) Cu(1)…Сu(1)ⅱ 0.261 1(1) Сu(1)-I(1)ⅰ 0.273 4(1) Cu(2)-N(2)ⅲ 0.180 5(4) Cu(2)-N(3)ⅳ 0.180 8(4) Cu(2)-I(1) 0.298 8(1) I(1)-Cu(1)ⅲ 0.259 9(1) І(1)-Cu(1)ⅰ 0.273 4(1) N(1)-Cu(1)-I(1) 119.6(1) N(1)-Сu(1)-Сu(1)ⅰ 137.22(11) I(1)-Сu(1)-Сu(1)ⅰ 65.76(3) N(1)-Сu(1)-I(1) 103.8(1) I(1)-Сu(1)-I(1)ⅱ 115.26(3) Сu(1)і-Сu1-I(1)ⅱ 111.2(1) N(1)-Cu(1)-Cu(1)ⅲ 161.81(1) I(1)-Сu(1)-Сu(1)ⅲ 61.19(2) Сu(1) i-Сu1-Сu(1)ⅲ 60.83(1) I(1)ⅱ-Сu(1)-Сu(1)ⅲ 63.32(3) N(1)-Cu(1)-Cu(1)ⅱ 127.33(14) І(1)-Сu(1)-Сu(1)ⅱ 112.55(3) N(1)-Сu(1)-I(1)ⅰ 90.8(1) I(1)-Сu(1)-I(1)ⅰ 116.74(3) Сu(1)ⅰ-Сu(1)-I(1)ⅰ 56.18(3) I(1)ⅱ-Сu(1)-I(1)ⅰ 107.43(2) Сu(1)ⅲ-Сu(1)-І(1)ⅰ 105.09(2) Сu(1)ⅱ-Сu(1)-I(1)ⅰ 58.13(2) N(2)ⅲ-Cu(2)-N(3)ⅳ 163.8(2) N(2)ⅲ-Cu(2)-I(1) 96.10(15) N(3)ⅳ-Cu(2)-I(1) 95.79(15) Cu(1)-I(1)-Cu(1)ⅲ 61.67(3) Сu(1)-I(1)-Сu(1)ⅰ 58.06(3) Сu(1)ⅲ-I(1)-Сu(1)ⅰ 58.55(3) Cu(1)-I(1)-Cu(2) 75.49(2) Сu(1)ⅲ-I(1)-Сu(2) 69.93(2) Сu(1)ⅰ-I(1)-Сu(2) 121.86(2) Symmetry codes: ⅰ -x+1, -y, -z+1; ⅱ -x+1, y+1/2, -z+1/2; ⅲ -x+1, y-1/2, -z+1/2 for 1; ⅰ -x+2, -y+1/2, z; ⅱ -y+5/4, x-3/4, -z+9/4; ⅲ y+3/4, -x+5/4, -z+9/4; ⅳ -x+2, -y+1, -z+2 for 2. CCDC: 1437794, 1; 1437795, 2.
2 Results and discussion
2.1 Syntheses of the complexes
The aadmtrz ligand in complex 1 is in situ prepared from the condensation reaction of acetonitrile and 3, 5-dimethyl-4-amino-1, 2, 4-triazole. This conden-sation reaction is observed between amino compounds and nitrile compounds under solvothermal conditions. The formation mechanism of in situ alkylation is suggested as follows: under high temperature and autogenous pressure, the hydrolysis breaks CH3CN molecules into CH3CONH2; the nucleophilic substitu-tion reaction of dmatrz and CH3CONH2 can generate aadmtrz, which attacks the nitrogen atom of dmatrz to form the 1-aminoethylidene group. The ligand in complex 2, dptrz, is from the reductive deaminization of the 3, 5-propyl-4-amino-1, 2, 4-triazole ligand. The cuprous iodide in both reactions serves not only as the metal catalyst for the other triazole ligands, but also as the source of the cuprous-iodide aggregates.
2.2 Structure description
2.2.1 Structure of {[Cu2(aadmtrz)I2]·CH3CN}n (1)
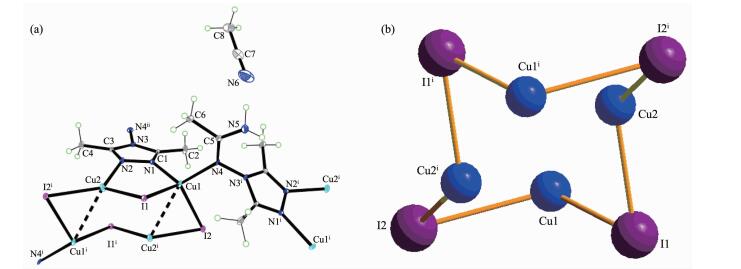
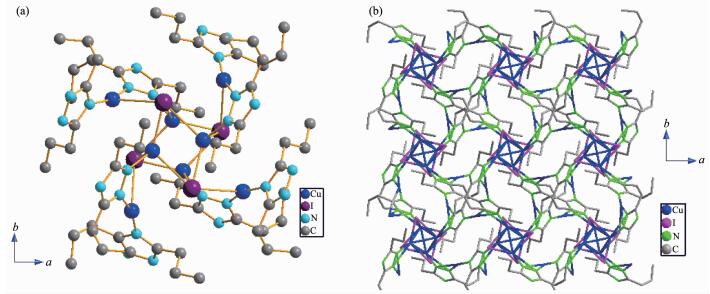
Complex 1 is a 2D coordination polymer with the monoclinic space group P21/c, and the asymmetric unit consists of two crystallographically independent cuprous ions, one aadmtrz group, two iodides, one CH3CN solvent molecule. As shown in Fig. 1a, its monomer is composed of a Cu4I4 cluster and two tridentate aadmtrz molecules. The Cu(Ⅰ) atoms are precisely coplanar, thus forming a parallelogram. Cu(1) shows tetrahedral coordination geometries [CuN2I2] by two I ions (I(1), I(2)) and two nitrogen atoms (N(1), N(4)) from the different addmtrz ligands. The distance of Cu(1)-N(1) (0.201 2(3) nm) is longer than that of Cu(1)-N(4) (0.206 3(3) nm), meanwhile the distance of Cu(1)-I(1) (0.268 9(1) nm) is also longer than that of Cu(1)-I(2) (0.267 1(1) nm). Cu(2) is in a slightly distorted [CuNI2] triangular planar site with one μ2-I(1) and two nitrogen atoms from different aadmtrz groups (N(2)ⅲ, N(3)ⅳ, Symmetry codes: ⅲ y+1/4, -x+7/4, -z+3/4; ⅳ -x+3/2, -y+3/2, -z+1/2). The dista-nces of Cu(2)-N (0.180 5(4) and 0.180 8(4) nm) are shorter than those of Cu(1)-N, and the distances of Cu(2)-I (0.257 8(1) and 0.252 2(1) nm) are shorter than that of Cu(1)-I (0.268 9(1) nm). The Cu(1)…Cu(2) interactions (0.288 4(1) nm) are crucial to build the Cu4I4 aggregates. The parallelogram Cu4I4 unit (Fig. 1b) has an inversion center and is formed by four iodine atoms bridges four copper atoms in the μ2-bridging modes, generating an 8-membered ring that has the chair-chair conformation of cyclooctane. This chair-chair conformation is similar to those similar complexes such as [Cu2I2(aadtz)]n[20].
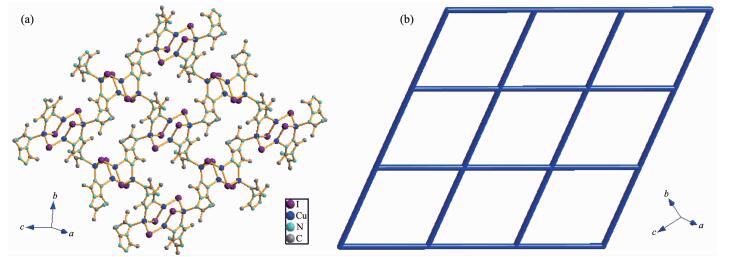
In this complex, each aadmtrz ligand bridges two Cu(Ⅰ) atoms on the longer side of the Cu parallelo-gram through N(1) and N(2) nitrogen donors, while 4-substituent N(4) and N(5) atoms link an adjacent Cu4I4 unit to form a 2D network (Fig. 2a). The overall structure of 1 can thus be simplified as a 2D (4, 4) rhombic-grid network with 4-connecting node of the Cu4I4 aggregates (Fig. 2b). The coplanar sheets of 1 are further built into a 3D supramolecular architecture through the hydrogen bonding interactions (Table 3).
D-H…A d(D-H) / nm d(Н…A)/nm d(D …A) / nm ∠D-H …A/(°) 1 N(5) -H(5) A...N(6) 0.091 0.252 0.326 4(7) 139 N(5)-H(5) B...I(1)ⅳ 0.091 0.293 0.373 7(4) 149 С(2)-Н(2)В...I(1)ⅱ 0.098 0.321 0.417 7(4) 169 C(4)-H(4)А...I(2)ⅰ 0.098 0.326 0.417 2(4) 155 C(6)-H(6) C...I(1)ⅴ 0.098 0.314 0.390 0(4) 135 С(8)-Н(8)А...І(1)ⅵ 0.098 0.326 0.399 7(5) 134 2 C(2)-H(2) A …I(1) 0.097 0.296 0.385 5(5) 153 С(2)-Н(2)В...І(1)ⅴ 0.097 0.298 0.392 5(5) 164 С(6) -Н(6)В…І(1)ⅳ 0.097 0.312 0.393 8(6) 143 С(2)-Н(2)А...І(1) 0.097 0.296 0.385 5(5) 153 С(2) -Н(2)В...І(1)ⅴ 0.097 0.298 0.392 5(5) 164 С(6)-Н(6)В...І(1)ⅳ 0.097 0.312 0.393 8(6) 143 Symmetry codes: ⅰ -x+1, -y, -z+1; ⅱ -x+1, y+1/2, -z+1/2; ⅳ x, -y-1/2, z-1/2; ⅴ -x+2, -y, -z+1; vi -x+2, y+1/2, -z+1/2; vii -x+1, -y, -z for 1; ⅳ -x+3/2, -y+3/2, -z+1/2; ⅴ y-1/4, -x+7/4, z-1/4 for 2. Furthermore, complex 1 is different from previously reported [Cu4I4(C4H8N4)4][15] with the similar reaction, where the ligand 3, 5-dimethyl-4-amino-1, 2, 4-triazole took part in the compound directly. This case is different from 3, 5-diethyl-4-amino-1, 2, 4-triazole compounds[20], where may attribute to the shorter -CH2 in the 3, 5-position of 1, 2, 4-triazole.
2.2.2 Structure of [Cu2(dptrz)I]n (2)
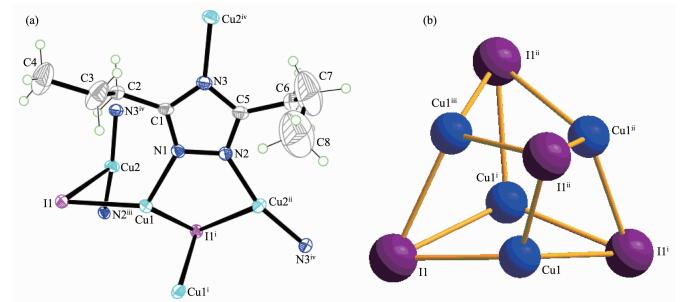
Complex 2 shows an interesting 3D porous structure with the tetragonal space group I41/a. As shown in Fig. 3a, the asymmetric unit of complex 2 contains two crystallographically independent Cu(Ⅰ) ions, one dptrz ligand, one iodide anions. Each Cu(1) or I(1) links three neighboring I(1) or Cu(1) to form a distorted Cu4I4 cubane unit (Fig. 3b), which is similar to those reported Cu4I4 tetramer units[24-25]. Each Cu(1) is tetrahedrally coordinated [CuNI3] by three I ions and a nitrogen from the bridging dptrz ligand. The distances of Cu(1)-I, Cu(1)-N and Cu…Cu are 0.249 1(1)~0.273 4(1), 0.191 4(4) and 0.254 4(1)~0.261 1(1) nm, respectively, which are shorter than those of isostruc-tural complex [Cu2(dtz)I][20], due to the longer -CH2 group in the ligand of 2. Furthermore, these values are comparable to those found in other cubane-like Cu4I4L4 complexes (L=nitrogen-containing ligand)[26]. Cu(2) is in a slightly distorted [CuN2I] triangular planar site with one μ3-I(1) and two nitrogen atoms from different dptrz groups (N(2)ⅲ, N(3)ⅳ, Symmetry codes: ⅲ y+1/4, -x+7/4, -z+3/4; ⅳ -x+3/2, -y+3/2, -z +1/2). The distances of Cu(2)-N (0.180 5(4), 0.180 8(4) nm) are shorter than those of Cu(1)-N, and the distance of Cu(2)-I is 0.298 7(7) nm, which is longer than that of Cu(1)-I. The angle of N(2)ⅲ-Cu(2)-N(3)ⅳ is 163.7(1)°.
Cu(2) atoms are bridged by μ1, 2-dptrz ligands forming single-stranded helices, with the helices being left-and right-handed along the fourfold axis (Cu4I4 unit locates on the fourfold inversion axes), respectively (Fig. 4a). Each helical chain along the c axis is further linked through Cu(1) of the Cu4I4 unit, forming a non-interpenetrating 3D framework (Fig. 4b). Topologically, when the Cu4I4 SBU are depicted as four-connecting nodes, and dptrz ligands are regarded as three-connecting nodes, complex 2 can be defined as a (3, 4)-connected framework, which is similar to the isostructural [Cu2I(dtz)]n[20]. Interesting, PLATON calculations[27] show that the potential solvent area volume of 2 is estimated to be 0.982 3 nm3, 19.5% of the unit cell volume (5.039 6 nm3), which may hold 10 H2O molecules. This may indicate that the construction of the cage-like motifs is accomplished by the -(CH2)n spacers of 3, 5-dialkyl-1, 2, 4-triazole.
2.3 PXRD analysis
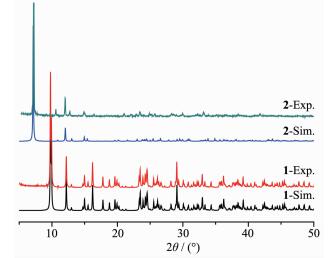
As shown in Fig. 5, the experiment PXRD patterns for 1 and 2 are in agreement with the simulated ones from the single-crystal X-ray data. The result indicates that the growth of crystals of both complexes is well-proportioned and consistent. The structures are representative of the bulk materials.
2.4 Luminescence properties
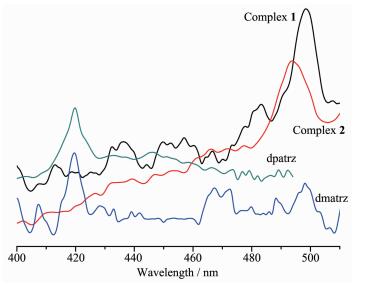
The emission spectra for 1 and 2 and the ligands dmatrz, dpatrz in the solid state at room temperature are shown in Fig. 6. The emission peaks of luminescence of ligands dmatrz, dpatrz are the same at 419 nm. It can be observed that complex 1 and 2 exhibit green emission bands at 498, 493 nm upon excitation at 266, 254 nm, respectively. The emission spectra for both complexes may be assigned, in chara-cter, to iodide(X)-to-ligand charge transfer (XLCT).
3 Conclusions
In summary, two cuprous halide coordination polymers containing Cu4I4 clusters layers as structural motifs have been successfully synthesized, which are further linked by 1, 2, 4-triazole to form extended 2D 4-connected grid frameworks (1) and 3D (3, 4)-connected (2) frameworks with 19.5% porosity. The Cu4I4 cluster in complex 1 is an inorganic 8-membered ring with a chair-chair conformation, while the one in complex 2 is a distorted cubane. The large spacers of alkyl side chains of 4-aminotriazole may accomplished the construction of the cage-like motifs. This provides a method to design the porous metal-organic frameworks (PMOFs). Both complexes exhibit similar green photo-luminescence at room temperature in the solid state.
Acknowledgements: The authors thank Dr. GAO Zeng-Qiang at line 3W1A of BSRF for his help in the single-crystal X-ray diffraction data collection and reduction.
Supporting information is available at http://www.wjhxxb.cn
-
-
[1]
Graham P M, Pike R D, Sabat M. et al. Inorg. Chem., 2000, 39:5121-5132 doi: 10.1021/ic0005341
-
[2]
Zhang Z Y, Deng Z P, Zhang X F, et al. CrystEngComm, 2014, 16:359-368 doi: 10.1039/C3CE41774C
-
[3]
Murdock C R, Jenkins D M. J. Am. Chem. Soc., 2014, 136:10983-10988 doi: 10.1021/ja5042226
-
[4]
Roesch P, Nitsch J, Lutz M, et al. Inorg. Chem., 2014, 53:9855-9859 doi: 10.1021/ic5014472
-
[5]
孙晓美, 宁为华, 刘建兰, 等.无机化学学报, 2013, 29(10):2171-2182 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?flag=1&file_no=20131024&journal_id=wjhxxbcnSUN Xiao-Mei, NING Wei-Hua, LIU Jian-Lan, et al. Chinese J. Inorg. Chem., 2013, 29(10):2171-2182 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?flag=1&file_no=20131024&journal_id=wjhxxbcn
-
[6]
Denysenko D, Grzywa M, Jelic J, et al. Angew. Chem. Int. Ed., 2014, 53:5832-5836 doi: 10.1002/anie.201310004
-
[7]
Li S L, Zhang X M. Inorg. Chem., 2014, 53:8376-8383 doi: 10.1021/ic500822w
-
[8]
Liu Z, Qayyum M F, Wu C, et al. J. Am. Chem. Soc., 2011, 133:3700-3703 doi: 10.1021/ja1065653
-
[9]
Jiang D P, Yao R X, Ji F, et al. Eur. J. Inorg. Chem., 2013, 2013:556-562 doi: 10.1002/ejic.201200949
-
[10]
Lee J Y, Kim H J, Jung J H, et al. J. Am. Chem. Soc., 2008, 130:13838-13839 doi: 10.1021/ja805337n
-
[11]
Lu J Y, Cabrera B R, Wang R J, et al. Inorg. Chem., 1999, 38:4608-4611 doi: 10.1021/ic990536p
-
[12]
Hou J J, Li S L, Li C R, et al. Dalton Trans., 2010, 39:2701-2707 doi: 10.1039/b922583h
-
[13]
Wu T, Li M, Li D, et al. Cryst. Growth Des., 2008, 8:568-574 doi: 10.1021/cg070639f
-
[14]
DeBord J R D, Lu Y J, Warren C J, et al. Chem. Commun., 1997, 15:1365-1366
-
[15]
Zhao Z G, Wu X Y, Zhang Q S, et al. Chin. J. Struct. Chem., 2010, 29(2):245-249
-
[16]
Hu M C, Wang Y, Zhai Q G, et al. Inorg. Chem., 2009, 48:1449-1468 doi: 10.1021/ic801574k
-
[17]
张俊珺, 阳年发, 张春华, 等.无机化学学报, 2010, 26(3):533-536 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?flag=1&file_no=20100328&journal_id=wjhxxbcnZHANG Jun-Jun, YANG Nian-Fa, ZHANG Chun-Hua, et al. Chinese J. Inorg. Chem., 2010, 26(3):533-536 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?flag=1&file_no=20100328&journal_id=wjhxxbcn
-
[18]
Ouellette W, Jones S, Zubieta J. CrystEngComm, 2011, 13:4457-4485 doi: 10.1039/c0ce00919a
-
[19]
Yan J Z, Lu L P, Su F, et al. Chin. J. Struct. Chem., 2015, 34(3):401-407
-
[20]
Zhao Z G, Yu R M, Wu X Y, et al. CrystEngComm, 2009, 11:2494-2499 doi: 10.1039/b821491c
-
[21]
Robert M H, James A G. J. Org. Chem., 1953, 18:872-877 doi: 10.1021/jo50013a016
-
[22]
Otwinowski Z, Minor W. Methods in Enzymology:Vol.276. Carter Jr C W, Sweet R M Ed. San Diego:Academic Press, 1997:307-326
-
[23]
Sheldrick G M. Acta Crystallogr. Sect. A, 2015, 71:3-8
-
[24]
Zhai Q G, Lu C Z, Chen S M, et al. Inorg. Chem. Commun., 2006, 9:819-822 doi: 10.1016/j.inoche.2006.04.033
-
[25]
Harvey P D, Knorr M M. Rapid Commun., 2010, 31:808-826 doi: 10.1002/marc.v31:9/10
-
[26]
Zhang D Q, Ding L, Xu W, et al. Chem. Lett., 2001:242-243
-
[27]
Spek A L. J. Appl. Crystallogr., 2003, 36:7-13 doi: 10.1107/S0021889802022112
-
[1]
-
Table 1. Crystallographic data and structure refinement for 1 and 2
1 2 Formula C8H14Cu2I2N6 C8H14Cu2N3I Formula weight 575.13 406.20 Crystal system Monoclinic Tetragonal Space group P21/c I41/a a / nm 0.954 7(1) 1.625 3(2) b/ nm 1.187 3(2) 1.625 3(2) c / nm 1.494 9(5) 1.907 8(4) β/(°) 111.23(3) V / nm3 1.579 5(7) 5.039 5(1) Crystal size / mm 0.45×0.40×0.40 0.05×0.04×0.03 Z 4 16 Dc / (g·cm-3) 2.419 2.142 μ/ mm-1 6.59 5.794 F(000) 1 072 3 104 θ range / (°) 2.9~26.0 2.51~25.05 Gocxlness-of-fit on F2 1.41 1.046 Reflection collected, unique 3 075, 3 067 2 234, 1 940 Rint 0.017 0.028 7 R1, wR2[I>2σ(I)] 0.019 1, 0.058 6 0.032 9, 0.089 2 R1, wR2(all data) 0.020 9, 0.072 5 0.038 6, 0.092 8 (Δ/σ)max 0.002 0.004 Largest diff peak and hole / (e • nm-3) 1 180, -1 580 697, -1 880 Table 2. Selected bond lengths (nm) and bond angles (°) of complexes 1 and 2
1 Сu(1)-N(1) 0.201 2(3) Cu(1)-N(4) 0.206 3(3) Cu(1)-I(1) 0.268 9(0) Сu(1)-I(2) 0.267 1(1) Cu(2)-N(2) 0.200 1(3) Cu(2)-I(1) 0.258 0(1) Сu(2)-I(2)ⅰ 0.252 3(1) Сu(1)...Сu(2)ⅰ 0.288 4(1) Сu(2)-Сu(1)ⅰ 0.288 4(1) Cu(2)-I(1)-Cu(1) 81.61(2) Сu(2)ⅰ-1(2)-Сu(1) 67.41(3) N(1)-Cu(1)-N(4) 120.04(1) N(1)-Cu(1)-I(2) 110.45(9) N(4)-Cu(1)-I(2) 103.08(8) N(1)-Cu(1)-I(1) 102.81(9) N(4)-Cu(1)-I(1) 104.16(9) I(2)-Cu(1)-I(1) 116.90(3) N(1)-Cu(1)-Cu(2)ⅰ 88.55(9) N(4)-Cu(1)-Cu(2)ⅰ 149.84(8) I(2)-Сu(1)-Сu(2)ⅰ 53.84(2) I(1)-Сu(1)-Сu(2)ⅰ 75.91(3) N(2)-Cu(2)-I(2)ⅰ 128.18(9) N(2)-Cu(2)-I(1) 105.90(8) I(2) i-Cu(2)-I(1) 122.09(3) N(2)-Cu(2)-Cu(1)ⅰ 121.26(9) I(2)ⅰ-Сu(2)-Сu(1)ⅰ 58.75(2) I(1)-Сu(2)-Сu(1)ⅰ 112.17(2) 2 Сu(1)-N(1) 0.191 5(4) Cu(1)-I(1) 0.249 1(1) Cu(1)…Сu(1)ⅰ 0.254 4(1) Сu(1)-І(1)ⅱ 0.259 9(1) Cu(1)…Cu(1)ⅲ 0.261 1(1) Cu(1)…Сu(1)ⅱ 0.261 1(1) Сu(1)-I(1)ⅰ 0.273 4(1) Cu(2)-N(2)ⅲ 0.180 5(4) Cu(2)-N(3)ⅳ 0.180 8(4) Cu(2)-I(1) 0.298 8(1) I(1)-Cu(1)ⅲ 0.259 9(1) І(1)-Cu(1)ⅰ 0.273 4(1) N(1)-Cu(1)-I(1) 119.6(1) N(1)-Сu(1)-Сu(1)ⅰ 137.22(11) I(1)-Сu(1)-Сu(1)ⅰ 65.76(3) N(1)-Сu(1)-I(1) 103.8(1) I(1)-Сu(1)-I(1)ⅱ 115.26(3) Сu(1)і-Сu1-I(1)ⅱ 111.2(1) N(1)-Cu(1)-Cu(1)ⅲ 161.81(1) I(1)-Сu(1)-Сu(1)ⅲ 61.19(2) Сu(1) i-Сu1-Сu(1)ⅲ 60.83(1) I(1)ⅱ-Сu(1)-Сu(1)ⅲ 63.32(3) N(1)-Cu(1)-Cu(1)ⅱ 127.33(14) І(1)-Сu(1)-Сu(1)ⅱ 112.55(3) N(1)-Сu(1)-I(1)ⅰ 90.8(1) I(1)-Сu(1)-I(1)ⅰ 116.74(3) Сu(1)ⅰ-Сu(1)-I(1)ⅰ 56.18(3) I(1)ⅱ-Сu(1)-I(1)ⅰ 107.43(2) Сu(1)ⅲ-Сu(1)-І(1)ⅰ 105.09(2) Сu(1)ⅱ-Сu(1)-I(1)ⅰ 58.13(2) N(2)ⅲ-Cu(2)-N(3)ⅳ 163.8(2) N(2)ⅲ-Cu(2)-I(1) 96.10(15) N(3)ⅳ-Cu(2)-I(1) 95.79(15) Cu(1)-I(1)-Cu(1)ⅲ 61.67(3) Сu(1)-I(1)-Сu(1)ⅰ 58.06(3) Сu(1)ⅲ-I(1)-Сu(1)ⅰ 58.55(3) Cu(1)-I(1)-Cu(2) 75.49(2) Сu(1)ⅲ-I(1)-Сu(2) 69.93(2) Сu(1)ⅰ-I(1)-Сu(2) 121.86(2) Symmetry codes: ⅰ -x+1, -y, -z+1; ⅱ -x+1, y+1/2, -z+1/2; ⅲ -x+1, y-1/2, -z+1/2 for 1; ⅰ -x+2, -y+1/2, z; ⅱ -y+5/4, x-3/4, -z+9/4; ⅲ y+3/4, -x+5/4, -z+9/4; ⅳ -x+2, -y+1, -z+2 for 2. Table 3. Hydrogen bonds geometry for 1 and 2
D-H…A d(D-H) / nm d(Н…A)/nm d(D …A) / nm ∠D-H …A/(°) 1 N(5) -H(5) A...N(6) 0.091 0.252 0.326 4(7) 139 N(5)-H(5) B...I(1)ⅳ 0.091 0.293 0.373 7(4) 149 С(2)-Н(2)В...I(1)ⅱ 0.098 0.321 0.417 7(4) 169 C(4)-H(4)А...I(2)ⅰ 0.098 0.326 0.417 2(4) 155 C(6)-H(6) C...I(1)ⅴ 0.098 0.314 0.390 0(4) 135 С(8)-Н(8)А...І(1)ⅵ 0.098 0.326 0.399 7(5) 134 2 C(2)-H(2) A …I(1) 0.097 0.296 0.385 5(5) 153 С(2)-Н(2)В...І(1)ⅴ 0.097 0.298 0.392 5(5) 164 С(6) -Н(6)В…І(1)ⅳ 0.097 0.312 0.393 8(6) 143 С(2)-Н(2)А...І(1) 0.097 0.296 0.385 5(5) 153 С(2) -Н(2)В...І(1)ⅴ 0.097 0.298 0.392 5(5) 164 С(6)-Н(6)В...І(1)ⅳ 0.097 0.312 0.393 8(6) 143 Symmetry codes: ⅰ -x+1, -y, -z+1; ⅱ -x+1, y+1/2, -z+1/2; ⅳ x, -y-1/2, z-1/2; ⅴ -x+2, -y, -z+1; vi -x+2, y+1/2, -z+1/2; vii -x+1, -y, -z for 1; ⅳ -x+3/2, -y+3/2, -z+1/2; ⅴ y-1/4, -x+7/4, z-1/4 for 2. -

 扫一扫看文章
扫一扫看文章
计量
- PDF下载量: 6
- 文章访问数: 1092
- HTML全文浏览量: 110


 下载:
下载:






 下载:
下载:

