
含有双膦配体的银(Ⅰ)配合物的合成、表征和荧光性质
-
关键词:
- 4, 5-双二苯基膦-9, 9-二甲基氧杂蒽
- / 双((2-二苯膦基)苯基)醚
- / 银
- / 荧光
English
Syntheses, Characterization and Luminescent Properties of Silver(Ⅰ) Complexes Based on Diphosphine Ligands
-
0 Introduction
In recent years, the study of silver(Ⅰ) complexes based on organic phosphine, especially diphosphine ligands, has attracted considerable attention[1-4]. In coor-dination chemistry, self-assembled silver(Ⅰ) complexes usually have structural diversity and potential appli-cations in luminescent material and antimicrobial[5-8]. According to the hard-soft-acid-base (HSAB) theory, as P-donor ligands, bis(2-(diphenylphosphino)phenyl)ether (DPEphos) and 9, 9-dimethyl-4, 5-bis(diphenyl-phosphanyl)xanthene (XANTphos) can both easily coordinate with Ag(Ⅰ) salts. Since the transition metal chemistry and catalytic function of DPEphos and XANTphos ligands were studied by van Leeuwan and his co-workers[9] for the first time, various researches about Ag(Ⅰ) complexes on the two ligands have been published[10-12].
Ag(Ⅰ)-DPEphos/XANTphos complexes are widely studied by us and other researchers[13-17]. In these complexes, DPEphos/XANTphos is a kind of chelated P-donor ligands, and some bridged co-ligands help to form 1D infinite chains. In some complexes the existence of weak interactions makes the 1D infinite chain structures expanding to 2D networks or 3D architectures. The type of weak interactions appeared in these complexes are hydrogen bond, C-H…π, π…π, anion…π and so on, which can also help to stabilize the supramolecular coordination polymers[8].
In this paper, two novel Ag-diphosphine comp-lexes, namely [Ag(XANTphos)]BF4 (1) and [Ag2Cl2(DPEphos)2]·2CH2Cl2 (2), have been synthesized and characterized by X-ray diffraction, IR, 1H NMR and fluorescence spectra. The weak interactions and lumin-escent properties of these complexes are discussed.
1 Experimental
1.1 Materials and measurements
All chemical reagents are commercially available and used without furthermore treatment. FT-IR spectra (KBr pellets) were measured on a Perkin-Elmer Infrared spectrometer. C, H and N elemental analysis were carried out on an Elementar Vario MICRO CUBE (Germany) elemental analyzer. Room-temperature fluorescence spectra were measured on F-4500 FL Spectrophotometer. 1H NMR was recorded at room temperature with a Bruker DPX 600 spectro-meter.
1.2 Synthesis of [Ag(XANTphos)]BF4 (1)
Complex 1 was prepared by the reaction of AgBF4 (0.038 4 g, 0.2 mmol), XANTphos (0.115 7 g, 0.2 mmol) and dmp (0.041 7 g, 0.2 mmol, dmp=neocuproine) in the mixed solvents of 5 mL CH2Cl2 and 5 mL CH3OH. The mixture was stirred for 6 h and filtered. Colorless crystals were obtained from the filtrate after standing at room temperature for several days. Yield: 71%. Element analysis Calcd. for C79H68AgBF4O3P4(%): C, 68.51; H, 4.92. Found(%): C, 68.08; H, 4.61. IR data (cm-1, KBr pellets): 3 514w, 3 053w, 2 958w, 1 971w, 1 815w, 1 621m, 1 590m, 1 502m, 1 478vs, 1 434vs, 1 402s, 1 360s, 1 327s, 1 306m, 1 284s, 1 218s, 1 198m, 1 151m, 1 057s, 870m, 857m, 801m, 778s, 746s, 610w, 588w, 548w, 537w, 514s, 498s, 460m. 1H NMR (600 MHz, CDCl3, 298 K): δ 7.9~8.4 (m, dissociative dmp-ph), 7.03~7.60 (m, overlap with the solvent peak signal, XANTphos-ph), 2.48 (s, dissociative dmp-CH3), 1.62 (s, 12H, XANTphos-CH3).
1.3 Synthesis of [Ag2Cl2(DPEphos)2]·2CH2Cl2 (2)
A mixture of AgCl (0.028 7 g, 0.02 mmol) and DPEphos (0.106 8 g, 0.2 mmol) was dissolved in a mixture of 5 mL CH2Cl2 and 5 mL CH3OH, stirred for 6 h and filtered. Colorless crystal 2 was obtained from the filtrate after standing at the room temperature for several days. Yield: 63%. Element analysis Calcd. for C74H60Ag2Cl6O2P4(%): C, 57.91; H, 3.91. Found(%): C, 57.56; H, 3.60. IR data (cm-1, KBr pellets): 3 422m, 3 051m, 1 624w, 1 586w, 1 564w, 1 479w, 1 461s, 1 434s, 1 259m, 1 222s, 1 159w, 1 095m, 1 027w, 876w, 801w, 745s, 695s, 507m, 422w.1H NMR (600 MHz, CDCl3, 298 K): δ 7.16~6.66 (m, CHbenzene), 5.29 (s, CHCH2Cl2)
1.4 Structure determination
Single crystals of the title complexes were mounted on a Bruker Smart 1000 CCD diffractometer equipped with a graphite-monochromated Mo Kα (λ=0.071 073 nm) radiation at 298 K. Semi-empirical absorption corrections were applied using SABABS program[18]. All the structures were solved by direct methods using SHELXS program of the SHELXTL-97 package and refined with SHELXL-97[19-20]. Metal atom centers were located from the E-maps and other non-hydrogen atoms were located in successive difference Fourier syntheses. The final refinements were performed by full matrix least-squares methods with anisotropic thermal parameters for non-hydrogen atoms on F2. The hydrogen atoms were generated geometrically and refined with displacement parameters riding on the concerned atoms.
Crystallographic data and experimental details for structural analysis are summarized in Table 1, and selected bond lengths and angles of complexes 1~2 are summarized in Table 2.
Complex 1 2 Formula C79H68AgBF4O3P4 C74H60Ag2Cl6O2P4 Formula weight 1 383.89 1 533.54 Crystal system Triclinic Triclinic Space group P1 P1 Crystal size / mm 0.45×0.40×0.30 0.36×0.328×0.31 a/nm 1.303 20(11) 1.202 80(11) b / nm 1.320 71(12) 1.258 11(12) c / nm 2.033 09(16) 1.422 69(13) α/(°) 95.033 0(10) 114.590(3) β/(°) 93.985 0(10) 98.963 0(10) γ/(°) 98.125(2) 107.292(2) V/ nm3 3.438 8(5) 1.769 7(3) Z 2 1 F(000) 1 428 776 Goodness-of-fit on F2 1.027 1.077 Rint 0.039 3 0.064 4 R1 [I > 2σ(I)]a 0.051 2 0.067 8 wR2[I > 2σ(I)]b 0.084 1 0.152 2 R1 (all data)a 0.098 6 0.123 9 wR2 (all data)b 0.092 4 0.166 0 a R=∑(||Fo|-|Fc||)/∑|Fo|; b wR=[∑w(|Fo|2-|Fc|2)2/∑w(Fo2)]1/2. Complex 1 Ag(1)-P(3) 0.257 9(4) Ag(1)-P(2) 0.259 1(8) Ag(1)-P(1) 0.258 5(6) Ag(1)-P(4) 0.256 0(5) P(1)-Ag(1)-P(2) 108.00(7) P(3)-Ag(1)-P(4) 104.86(9) P(2)-Ag(1)-P(4) 107.46(9) P(1)-Ag(1)-P(3) 106.95(8) Complex 2 Ag(1)-P(1) 0.246 4(2) Ag(1)-Cl(1) 0.262 6(2) Ag(1)-P(2) 0.249 5(2) Ag(1)-Cl(1)ⅰ 0.259 3(2) P(1)-Ag(1)-P(2) 111.33(7) P(1)-Ag(1)-Cl(1) 114.67(7) P(2)-Ag(1)-Cl(1) 105.05(8) P(1)-Ag(1)-Cl(1)ⅰ 116.76(8) P(2)-Ag(1)-Cl(1)ⅰ 112.59(7) Cl(1)-Ag(1)-Cl(1)ⅰ 94.92(6) Symmetry codes: ⅰ 1-x, 1-y, 1-z CCDC: 1555835, 1; 1508048, 2.
2 Results and discussion
2.1 Syntheses of the complexes
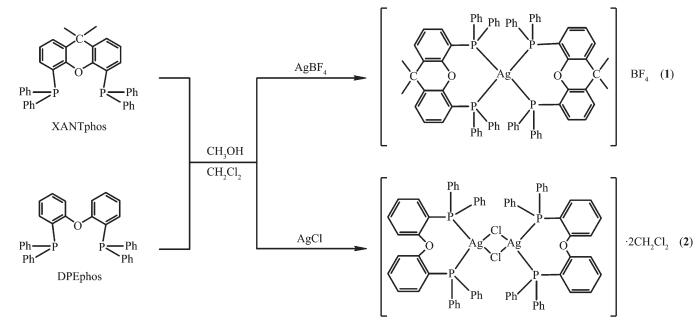
Two functional Ag(Ⅰ) complexes 1~2 have been synthesized by one-pot reaction of different silver(Ⅰ) salts with DPEphos and XANTphos ligands (Scheme 1). The influence of the anions on the coordination modes of the complexes has been discussed in the literatures[21-22]. We also have reported the structures of some metal complexes which are affected by the coordination modes[23-24]. In complex 2, two silver atoms are connected by halogen atoms; however, the tetrafluoroborate anion is free in complex 1.
2.2 Description of crystal structures
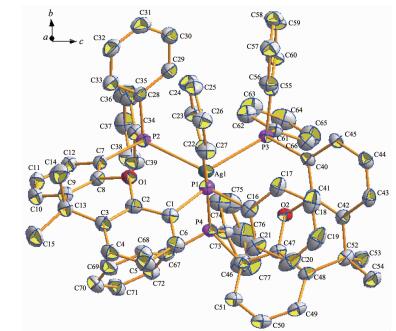
Complex 1 crystallizes in the triclinic crystal system and contains a crystallographic center of symmetry (Fig. 1). In complex 1, the central ion Ag(Ⅰ) forms a ring by the chelating bisphosphine ligand (XANTphos), and each Ag(Ⅰ) is coordinated with a P atom from a different XANTphos molecule to form a distorted tetrahedron. Compared to the similar Ag-XANTphos complex, the Ag-P (0.258 5 and 0.259 1 nm) distance in complex 1 is longer than that in [AgBr(XANTphos)(py2SH)]·C2H5OH (Ag-P 0.248 4 and 0.250 0 nm)[16], and the P-Ag-P angle (108.00°) is smaller than that in the complex above.
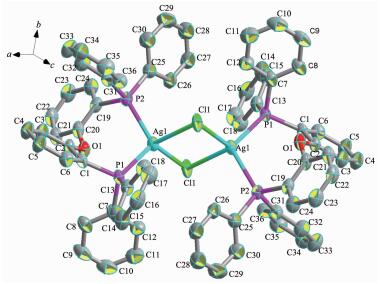
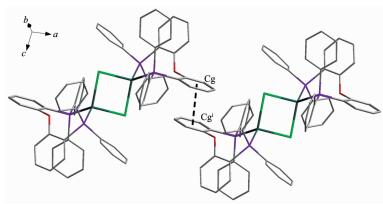
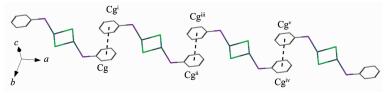
Complex 2 consists of inversion symmetric dimers with a diamond-shaped Ag2Cl2 group at the center. In the asymmetric unit of complex 2, each DPEphos ligand chelates one silver(Ⅰ) cation with a free dichlorome-thane in it. Two silver cations are connected by two Cl- anions in the formation of a binuclear structure (Fig. 2). The sum of the internal angles of the four-member ring [-Ag-Cl-Ag-Cl-] in the Ag2Cl2 core are 360°, demonstrating that the ring is a parallelogram. The geometry around each silver center is distorted tetrahedral, which is evident from the angles around the Ag ion in the range of 94.92°~116.76°. The distance of Ag…Ag is 0.352 9 nm, which exceeds the sum of two van der Waals radius of silver atoms (0.344 0 nm)[25], so the metal-metal interaction can be neglected in complex 2. The bite angle of the DPEphos ligand is 111°, which is 9° larger than its natural bite angle (βn)[9]. In both of two complexes in this paper, the ether O atoms of the DPEphos ligands are always at a nonbonding distance from the metal centers. The lattice structure is stabilized by an inter-molecular π…π stacking interactions (Fig. 3) between the C19-C24 rings of DPEphos ligand, with the interplanar distance of 0.381 3 nm. The center-of-mass coordinate of the ring is (-0.014 48, 0.389 77, 0.561 57) and the dihedral angle is 0°. The adjacent molecules in complex 2 are connected by the π…π weak interactions to form a 1D infinite chain (Fig. 4). The distance of Cl…π weak interaction (0.383 0 nm) found in complex 2 is shorter than that in the litera-ture[8]. Those weaker interactions stabilize the supra-molecular frameworks.
2.3 Infrared and 1H NMR spectroscopy
The infrared spectra of 1~2 show C-C stretch vibration of the phenyl rings whose absorptions are found in 1 434 cm-1. The middle absorptions around 3 051 cm-1 are caused by C-H vibration of the phenyl rings.The absorption in 1 057 cm-1 is derived from tetrafluoroborate (BF4-) in complex 1.
The singlet near δ 1.62 in 1H NMR spectrum of 1 is assigned to the protons of methylene groups in the aromatic rings from XANTphos ligand. The 1H NMR spectrum of complexes 2 exhibit signals (multiple peaks) between 7.1 and 6.6, which can be attributed to protons from the benzene rings of the bis(2-(diphenylphosphino)phenyl)ether (DPEphos) ligand. In complex 2, the signals of protons from the dichloro-methane solvent molecule are shown at 5.29.
2.4 Fluorescence spectra
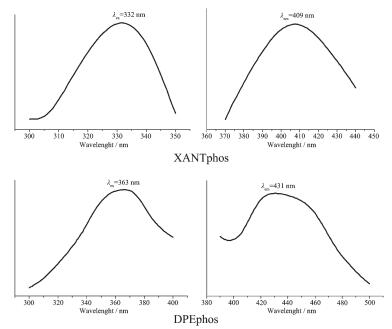
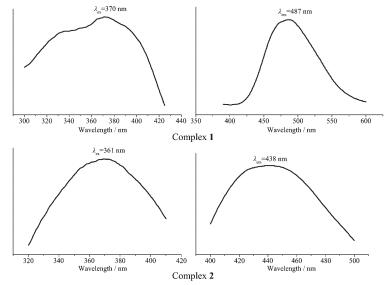
At room temperature, the solid-state excitation and emission spectra of complexes 1~2 and the DPEphos and XANTphos ligands were measured. When being excited at 332 nm, the XANTphos ligand displays a fluorescence emission peak at 409 nm, and the emission peak of the DPEphos ligand is found at 431 nm with excitation at 363 nm (Fig. 5). It was found that the emission peak is centered at 487 nm with λex=370 nm for complex 1, and it is centered at 438 nm with λex=361 nm for complex 2 (Fig. 6). Compared to the free ligand, the emission peak of 1 is red-shifted by about 78 nm, however, the emission peak of 2 is red-shifted by about 7 nm. The difference of luminescent properties is attributed to their different structures. These shifts of emission peak are derived from ligand-centered π-π* transition.
3 Conclusions
Two novel Ag-biphosphine complexes, namely [Ag(XANTphos)]BF4 and [Ag2Cl2(DPEphos)2]·2CH2Cl2, have been synthesized and characterized by X-ray diffraction, IR, 1H NMR and fluorescence spectra. Single-crystal X-ray diffraction analysis reveals that the complex 1 crystallizes with triclinic system, P1 space group. Complex 2 consists of inversion symmetric dimers with a diamond-shaped Ag2Cl2 group at the center. The adjacent molecules in complex 2 are connected by π…π weak interactions to form a 1D infinite chain. In the emission spectra, shifts of emission peak are derived from ligand-centered π-π* transition. We hope our results could offer a new strategy for the design of supramolecular complexes.
-
-
[1]
Cook A W, Nguyen T D, Buratto W R, et al. Inorg. Chem., 2016, 55:12435-12440 doi: 10.1021/acs.inorgchem.6b02385
-
[2]
Streitberger M, Schmied A, Hoy R, et al. Dalton Trans., 2016, 45:11644-11649 doi: 10.1039/C6DT00945J
-
[3]
Zavras A, Khairallah G N, Connell T U, et al. Inorg. Chem., 2014, 53:7429-7437 doi: 10.1021/ic5007499
-
[4]
李中锋, 张彦茹, 崔洋哲, 等.无机化学学报, 2015, 31(8):1637-1643 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20150824&flag=1LI Zhong-Feng, ZHANG Yan-Ru, CUI Yang-Zhe, et al. Chinese J. Inorg. Chem., 2015, 31(8):1637-1643 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20150824&flag=1
-
[5]
Najafabadia B K, Corrigan J F. Chem. Commun., 2015, 51:665-667 doi: 10.1039/C4CC06560C
-
[6]
Lim S H, Cohen S M. Inorg. Chem., 2013, 52:7862-7872 doi: 10.1021/ic302840x
-
[7]
Jaros S W, Silva M C, Florek M, et al. Inorg. Chem., 2016, 55:5886-5894 doi: 10.1021/acs.inorgchem.6b00186
-
[8]
Rajput G, Yadav M K, Drew M G B, et al. Inorg. Chem., 2015, 54:2572-2579 doi: 10.1021/ic502688h
-
[9]
Kranenburg M, van der Burgt Y, Kamer P, et al. Organome-tallics, 1995, 14:3081-3089 doi: 10.1021/om00006a057
-
[10]
Kaeser A, Moudam O, Accorsi G, et al. Eur. J. Inorg. Chem., 2014:1345-1355
-
[11]
Freudenmann D, Feldmann C. Inorg. Chim. Acta, 2011, 375:311-313 doi: 10.1016/j.ica.2011.05.006
-
[12]
Moudam O, Tsipis A C, Kommanaboyina S, et al. RSC Adv., 2015, 5:95047-95053 doi: 10.1039/C5RA12921D
-
[13]
Gao S, Li Z F, Liu M, et al. Polyhedron, 2014, 83:10-15 doi: 10.1016/j.poly.2014.03.028
-
[14]
Cui Y Z, Yuan Y, Han H L, et al. Z. Anorg. Allg. Chem., 2016, 642(18):953-959 doi: 10.1002/zaac.201600193
-
[15]
Zhang Y R, Cui Y Z, Jin Q H, et al. Polyhedron, 2017, 122:86-98 doi: 10.1016/j.poly.2016.10.033
-
[16]
Kaltzoglou A, Cox P J, Aslanidis P. Polyhedron, 2007, 26:1634-1642 doi: 10.1016/j.poly.2006.12.006
-
[17]
张彦茹, 王梦秦, 崔洋哲, 等.无机化学学报, 2015, 31(10):2089-2094 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20151024&flag=1ZHANG Yan-Ru, WANG Meng-Qin, CUI Yang-Zhe, et al. Chinese J. Inorg. Chem., 2015, 31(10):2089-2094 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20151024&flag=1
-
[18]
SMART and SAINT, Siemens Analytical X-ray Instrument Inc., Madison, WI, 1996.
-
[19]
Sheldrick G M. SHELXS-97 and SHELXL-97, Madison, WI, USA, 1997.
-
[20]
Sheldrick G M. SHELXTL NT Ver. 5. 1, University of Göttingen, Germany, 1997.
-
[21]
Li X X, Gong Y Q, Zhao H X, et al. Inorg. Chem., 2014, 53:12127-12134 doi: 10.1021/ic501978u
-
[22]
Lee J, Kang Y, Cho N S, et al. Cryst.Growth Des., 2016, 16:996-1004 doi: 10.1021/acs.cgd.5b01544
-
[23]
Song L L, Jin Q H, Cui L N, et al. Inorg. Chim. Acta, 2010, 363:2425-2429 doi: 10.1016/j.ica.2010.03.075
-
[24]
Li J B, Fan W W, Liu M, et al. J. Mol. Struct., 2015, 1085:235-241 doi: 10.1016/j.molstruc.2014.12.068
-
[25]
Sculfort S, Braunstein P. Chem. Soc. Rev., 2011, 40:2741-2760 doi: 10.1039/c0cs00102c
-
[1]
-
Table 1. Crystallographic data for complexes 1~2
Complex 1 2 Formula C79H68AgBF4O3P4 C74H60Ag2Cl6O2P4 Formula weight 1 383.89 1 533.54 Crystal system Triclinic Triclinic Space group P1 P1 Crystal size / mm 0.45×0.40×0.30 0.36×0.328×0.31 a/nm 1.303 20(11) 1.202 80(11) b / nm 1.320 71(12) 1.258 11(12) c / nm 2.033 09(16) 1.422 69(13) α/(°) 95.033 0(10) 114.590(3) β/(°) 93.985 0(10) 98.963 0(10) γ/(°) 98.125(2) 107.292(2) V/ nm3 3.438 8(5) 1.769 7(3) Z 2 1 F(000) 1 428 776 Goodness-of-fit on F2 1.027 1.077 Rint 0.039 3 0.064 4 R1 [I > 2σ(I)]a 0.051 2 0.067 8 wR2[I > 2σ(I)]b 0.084 1 0.152 2 R1 (all data)a 0.098 6 0.123 9 wR2 (all data)b 0.092 4 0.166 0 a R=∑(||Fo|-|Fc||)/∑|Fo|; b wR=[∑w(|Fo|2-|Fc|2)2/∑w(Fo2)]1/2. Table 2. Selected bond distances (nm) and bond angles (°) for complexes 1~2
Complex 1 Ag(1)-P(3) 0.257 9(4) Ag(1)-P(2) 0.259 1(8) Ag(1)-P(1) 0.258 5(6) Ag(1)-P(4) 0.256 0(5) P(1)-Ag(1)-P(2) 108.00(7) P(3)-Ag(1)-P(4) 104.86(9) P(2)-Ag(1)-P(4) 107.46(9) P(1)-Ag(1)-P(3) 106.95(8) Complex 2 Ag(1)-P(1) 0.246 4(2) Ag(1)-Cl(1) 0.262 6(2) Ag(1)-P(2) 0.249 5(2) Ag(1)-Cl(1)ⅰ 0.259 3(2) P(1)-Ag(1)-P(2) 111.33(7) P(1)-Ag(1)-Cl(1) 114.67(7) P(2)-Ag(1)-Cl(1) 105.05(8) P(1)-Ag(1)-Cl(1)ⅰ 116.76(8) P(2)-Ag(1)-Cl(1)ⅰ 112.59(7) Cl(1)-Ag(1)-Cl(1)ⅰ 94.92(6) Symmetry codes: ⅰ 1-x, 1-y, 1-z -

 扫一扫看文章
扫一扫看文章
计量
- PDF下载量: 3
- 文章访问数: 2108
- HTML全文浏览量: 138


 下载:
下载:






 下载:
下载:

