 图Scheme 1
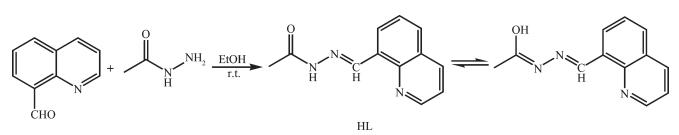
Synthesis route of HL
Scheme1.
Synthesis route of HL
图Scheme 1
Synthesis route of HL
Scheme1.
Synthesis route of HL

喹啉-8-甲醛乙酰腙锌/镉配合物的晶体结构及荧光性质
English
Crystal Structures and Fluorescence Property of Zn(Ⅱ)/Cd(Ⅱ) Complexes Based on N-((quinolin-8-yl)methylene)acetohydrazide
-
Key words:
- acylhydrazone
- / Zn(Ⅱ) complex
- / Cd(Ⅱ) complex
- / quinoline
- / fluorescence
-
Schiff bases are an important class of ligands in coordination chemistry and have been found extensive application in different fields[1-2]. As one of the most promising systems, the relevant semicarbazones and thiosemicarbazones involve condensed heterocycle, especially quinoline, have been paid much attention due to their potentially biological activities[3-6]. However, acylhydrazones, as their structurally analogous, have been paid much less attention[7-8]. Recently, several quinoline based acylhydrazone chemosensors for the fluorescent detection of metal ions have been reported in the literature, most of which function by coordina-tion reaction with ions[9-11]. Nevertheless, the crystal structures of corresponding complexes are relatively scarce[11].
Our previous work also shows that the acylhy-drazone ligand HL (Scheme 1), namely N-((quinolin-8-yl)methylene)acetohydrazide is an excellent fluorescent probe for the detection for Zn(Ⅱ) ions[11]. Therefore, in this paper, three Zn(Ⅱ) and Cd(Ⅱ) complexes with HL have been synthesized and structural determined by single-crystal X-ray diffraction. In addition, the fluorescence properties of the complexes in CH3CN solution were investigated.
1 Experimental
1.1 Materials and measurements
Solvents and starting materials for synthesis were purchased commercially and used as received. Elemental analysis was carried out on an Elemental Vario EL analyzer. The IR spectra (ν=4 000~400 cm-1) were determined by the KBr pressed disc method on a Bruker V70 FT-IR spectrophotometer. The UV spectra were recorded on a Purkinje General TU-1800 spectrophotometer. Fluorescence spectra were deter-mined on a Varian CARY Eclipse spectrophotometer, in the measurements of emission and excitation spectra the pass width is 5 nm.
1.2 Preparations of complexes 1~3
As shown in Scheme 1, the ligand HL was produced by condensation of 8-formylquinoline and acethydrazide in ethanol at room temperature accor-ding to the literature method[11]. The complexes 1~3 were generated by reaction of the ligand HL (5 mmol) with equimolar of ZnSO4, CdCl2 and CdI2 in methanol solution (10 mL) at room temperature for 1 h, respe-ctively. Crystals suitable for X-ray diffraction analysis were obtained by evaporating the corresponding reaction solutions at room temperature.
1: Colorless plates. Anal. Calcd. for C12H15N3O7SZn (%): C: 35.09; H: 3.68; N: 10.23. Found(%): C: 34.75; H: 3.85; N: 9.94. FT-IR (cm-1): ν(C=O) 1 655, ν(C=N) 1 592, ν(C=N)pyrazine 1 560.
2: Colorless blocks. Anal. Calcd. For C12H11N3O Cl2Cd(%):C: 36.35; H: 2.80; N: 10.60. Found (%): C: 36.42; H: 3.05; N: 10.37. FT-IR (cm-1): ν(C=O) 1 654, ν(C=N) 1 590, ν(C=N)pyrazine 1 558.
3: Colorless blocks. Anal. Calcd. For C12H11N3OI2 Cd(%): C: 24.87; H: 1.91; N: 7.25. Found(%): C: 25.00; H: 2.18; N: 7.02. FT-IR (cm-1): ν(C=O) 1 646, ν(C=N) 1 586, ν(C=N)pyrazine 1 555.
1.3 X-ray crystallography
The X-ray diffraction measurement for complexes 1~3 were performed on a Bruker SMART APEX Ⅱ CCD diffractometer equipped with a graphite mono-chromatized Mo Kα radiation (λ=0.071 073 nm) by using φ-ω scan mode at 296(2) K. Semi-empirical absorption correction was applied to the intensity data using the SADABS program[12]. The structures were solved by direct methods and refined by full matrix least-square on F2 using the SHELX-97 program[13]. All non-hydrogen atoms were refined anisotropically. All the H atoms were positioned geometrically and refined using a riding model. Details of the crystal parameters, data collection and refinements for complexes 1~3 are summarized in Table 1.
1 2 3 Empirical formula C12H15N3O7SZn C12H11N3OCl2Cd C12H11N3OI2Cd Formula weight 410.70 396.54 579.44 Size / mm 0.15×0.12×0.06 0.20×0.18×0.16 0.18×0.16×0.15 Crystal system Monoclinic Monoclinic Monoclinic Space group P21/n P21/c P21/n a / nm 1.033 9(7) 0.792 90(19) 0.746 79(12) b / nm 0.664 5(4) 1.523 4(3) 1.659 7(2) c / nm 2.179 5(14) 1.404 0(3) 1.263 00(12) β/(°) 93.915(11) 123.322(10) 90.132 0(10) V/ nm3 1.493 8(16) 1.417 1(5) 1.565 4(3) Z 4 4 4 Dc / (g·cm-3) 1.826 1.859 2.459 Unique 2 616 2 495 2 756 Rint 0.072 7 0.021 1 0.022 0 GOF 1.032 1.070 1.053 R indices [I/>2σ(I)] R1=0.044 3, wR2=0.079 5 R1=0.026 0, wR2=0.063 5 R1=0.026 9, wR2=0.059 7 R indices (all data) R1=0.091 6, wR2=0.092 7 R1=0.031 4, wR2=0.066 8 R1=0.032 7, wR2=0.062 5 CCDC: 1562151, 1; 1562152, 2; 1562153, 3.
2 Results and discussion
2.1 Crystal structures description
The diamond drawings of complexes 1~3 are shown in Fig. 1. Selected bond distances and angles are listed in Table 2. As shown in Fig. 1a, 1 contains one discrete cationic Zn(Ⅱ) complex and one crystal water molecule in the asymmetric unit. The center Zn(Ⅱ) ion with a distorted octahedron geometry is coordinated by one neutral hydrazone with ONN donor set, one coordinated water molecule and two O atoms from two independent μ2-bridged sulfate anions, thus forming one dimension chain-like framework along b axis. In addition, in the solid state, the chains were further linked into a 2D supramolecular network by inter-molecular N-H…O and O-H…O hydrogen bonds (Fig. 1d and Table 3).
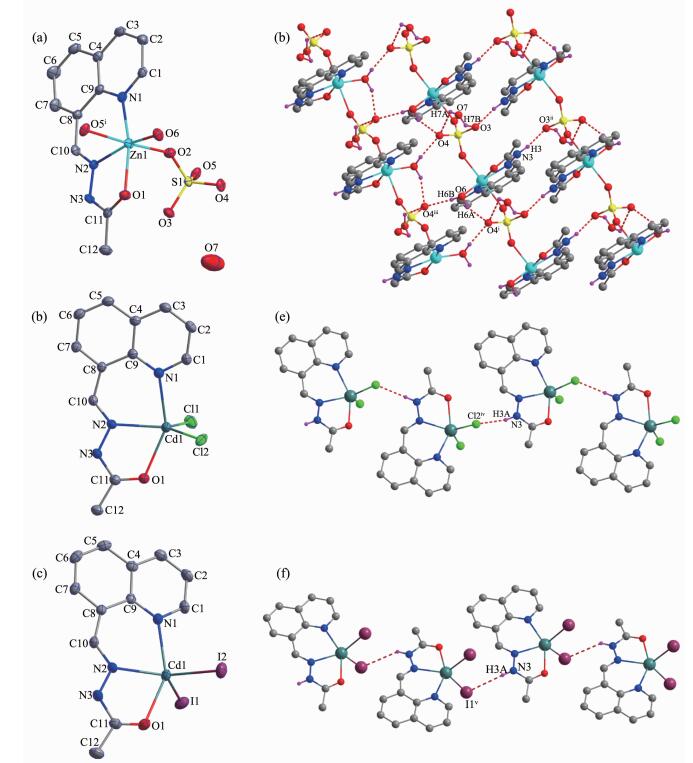 图1
Diamond drawing of 1~3 (a~c) with 30% thermal ellipsoids; Extended 2D supramolecular structure in complex 1 (d); Chain-like structures in complex 2 (e, along c axis) and 3 (f) formed by hydrogen bonds (shown in dashed line), respectively
Figure1.
Diamond drawing of 1~3 (a~c) with 30% thermal ellipsoids; Extended 2D supramolecular structure in complex 1 (d); Chain-like structures in complex 2 (e, along c axis) and 3 (f) formed by hydrogen bonds (shown in dashed line), respectively
图1
Diamond drawing of 1~3 (a~c) with 30% thermal ellipsoids; Extended 2D supramolecular structure in complex 1 (d); Chain-like structures in complex 2 (e, along c axis) and 3 (f) formed by hydrogen bonds (shown in dashed line), respectively
Figure1.
Diamond drawing of 1~3 (a~c) with 30% thermal ellipsoids; Extended 2D supramolecular structure in complex 1 (d); Chain-like structures in complex 2 (e, along c axis) and 3 (f) formed by hydrogen bonds (shown in dashed line), respectively
1 Zn1-N1 0.209 9(4) Zn1-N2 0.205 9(4) Zn1-O1 0.2l5 8(3) Zn1-O2 0.211 4(3) Zn1-O6 0.200 0(3) Zn1-O5ⅰ 0.227 9(3) O6-Zn1-N2 161.91(14) N1 -Zn1 -O2 87.73(14) O6-Zn1-O5ⅰ 86.08(12) O6-Zn1-N1 107.02(15) O6-Zn1 -O1 87.47(14) N2-Zn1-O5ⅰ 86.09(13) N2-Zn1-N1 88.97(15) N2-Zn1 -O1 76.09(15) N1-Zn1-O5ⅰ 87.98(13) O6-Zn1 -O2 95.30(13) N1 -Zn1 -O1 164.90(13) 02-Zn1-O5ⅰ 175.71(12) N2-Zn1 -O2 93.70(13) 02-Zn1 -O1 95.28(13) H-Zn1-O5ⅰ 88.83(12) 2 Cd1-N1 0.235 6(2) Cd1-N2 0.231 4(2) Cd1-O1 0.236 0(2) Cd1-Cl1 0.241 82(11) Cd1-Cl2 0.244 57(10) Cl1-Cd1-Cl2 116.27(4) N2-Cd1-N1 79.00(8) N2-Cd1-Cl1 117.11(7) N2-Cd1-Cl2 126.40(7) N2-Cd1-O1 69.66(8) N1-Cd1-Cl1 97.34(7) N1-Cd1-Cl2 98.92(7) N1-Cd1-O1 147.27(8) O1-Cd1-Cl1 105.00(6) O1-Cd1-Cl2 92.37(6) 3 Cd1-N1 0.237 8(4) Cd1-N2 0.230 8(3) Cd1-O1 0.238 4(3) Cd1-I1 0.274 72(7) Cd1-I2 0.273 21(5) I2-Cd1-I1 109.208(16) N2-Cd1-N1 78.88(12) N2-Cd1-I2 127.43(9) N2-Cd1-I1 122.37(9) N2-Cd1-O1 69.29(12) N1-Cd1-I2 100.14(9) N1-Cd1-I1 102.92(9) N1-Cd1-O1 148.15(12) O1-Cd1-I2 98.28(9) O1-Cd1-I1 95.17(10) Symmetry codes: ⅰ x, -1+y, z D-H …A d(D-H) / nm d(H …A) / nm d(D …A) / nm ∠DHA/(°) 1 O6-H6A …O4ⅰ 0.085 0.191 0.270 9(5) 155.1 N3-H3 …O3ⅱ O.O86 0.193 0.274 4(5) 158.2 O6-H6B …O4ⅲ 0.085 0.189 0.270 1(5) 159.6 O7-H7A …O4 0.085 0.26 0.317 4(6) 125.9 O7-H7B …O3 0.085 0.25 0.297 4(7) 116 2 N3-H3A …Cl2iv 0.086 0.235 0.318 9(3) 164.3 3 N3-H3A …I1v 0.086 0.311 0.383 1(4) 142.3 Symmetry codes: ⅰ x, -1+y, z; ⅱ 1-x, 1-y, -z; ⅲ 1.5-x, -0.5+y, 0.5-z; ⅳ x, 0.5-y, 0.5+z; ⅴ -0.5+x, 1.5-y, -0.5+z Similarly, the hydrazone HL acts as a neutral tridentate ligand in complexes 2 and 3 (Fig. 1b and 1c). Coordinated by two additional halide anions (chloride for 2, while iodide for 3), the Cd(Ⅱ) ion adopts a distorted square pyramid coordination geometry (τ=0.348 and 0.345 for complex 2 and 3, respectively)[7]. In the crystal, intermolecular N-H…Cl or N-H…I hydrogen bonds link the complex molecules of 2 or 3 into one dimension chains (Fig. 1e and 1f).
2.2 IR spectra
The FT-IR spectral region for both complexes is more or less similar due to the similar coordination modes of the ligands. The ν(C=O), ν(C=N)imine and ν(C=N)quinoline bands are at 1 673, 1 615 and 1 584 cm-1, respectively. They shift to lower frequency values in the complexes, indicating that the carbonyl O, imine N and quinoline N atoms take part in the coordina-tion[7-8, 14-15]. It is in accordance with the crystal structure study.
2.3 UV spectra
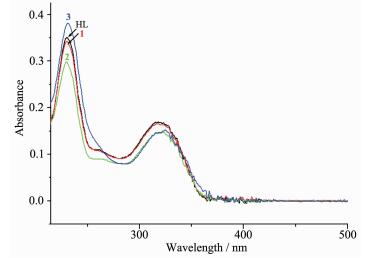
The UV spectra of the ligand HL, complexes 1~3 in CH3CN solution (c=1×10-5 mol·L-1) were measured at room temperature (Fig. 2). The spectra of HL features two main band located around 230 nm (ε=35 288 L·mol-1·cm-1) and 320 nm (ε=16 955 L·mol-1·cm-1), which could be assigned to characteristic π-π* transition of quinoline and imine units, respe-ctively[8]. Both bands have no shift while with absorption intensity change in the spectra of complexes 1~3 (ε1=34 327, 16 575 L·mol-1·cm-1; ε2=30 131, 14 854 L·mol-1·cm-1; ε3=38 244, 14 870 L·mol-1·cm-1). This fact supports the neutral mode of the ligand HL in three complexes[7].
2.4 Fluorescence spectra
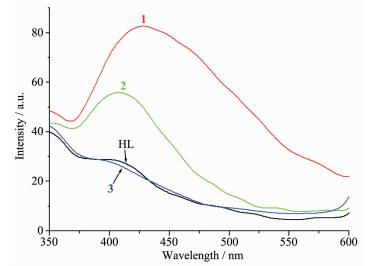
The fluorescence spectra of the ligand HL and complexes 1~3 have been studied in CH3CN solution (c=1×10-5 mol·L-1) at room temperature. The free Schiff base ligand HL exhibits almost none fluorescence emission when excited at 320 nm, primarily due to C=N isomerization. However, compl-exes 1 and 2 show remarkable peaks at about 428 and 408 nm under the same tested condition, respectively. Obviously, binding with Zn2+/Cd2+ inhibits the isome-rization of C=N, thereby increasing the fluorescence intensity through the CHEF mechanism[9-11]. In addition, it should be noted that complex 3 gives similar emission as the free ligand because of the heavy atom effect of the coordinated iodide anions.
-
-
[1]
Alagesan L, Bhuvanesh N S P, Dharmaraj N. Dalton Trans., 2013, 42:7210-7223 doi: 10.1039/c3dt50371b
-
[2]
Ye X P, Zhu T F, Wu W N, et al. Inorg. Chem. Commun., 2014, 47:60-62 doi: 10.1016/j.inoche.2014.07.022
-
[3]
Bourosh P N, Revenko M D, Stratulat E F, et al. Russ. J. Inorg. Chem., 2014, 59:545-557 doi: 10.1134/S0036023614060059
-
[4]
Revenko M D, Bourosh P N, Stratulat E F, et al. Russ. J. Inorg. Chem., 2010, 55:1387-1397 doi: 10.1134/S0036023610090093
-
[5]
毛盼东, 闫玲玲, 王文静, 等.无机化学学报, 2016, 32(3):555-560 doi: 10.11862/CJIC.2016.064MAO Pan-Dong, YAN Ling-Ling, WANG Wen-Jing, et al. Chinese J. Inorg. Chem., 2016, 32(3):555-560 doi: 10.11862/CJIC.2016.064
-
[6]
毛盼东, 韩学峰, 李珊珊, 等.无机化学学报, 2017, 33(4):692-698 doi: 10.11862/CJIC.2017.067MAO Pan-Dong, HAN Xue-Feng, LI Shan-Shan, et al. Chinese J. Inorg. Chem., 2017, 33(4):692-698 doi: 10.11862/CJIC.2017.067
-
[7]
李晓静, 吴伟娜, 徐周庆, 等.无机化学学报, 2015, 31(11):2265-2271 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20151123&flag=1LI Xiao-Jing, WU Wei-Na, XU Zhou-Qing, et al. Chinese J. Inorg. Chem., 2015, 31(11):2265-2271 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20151123&flag=1
-
[8]
常慧琴, 原知则, 赖晓晴, 等.无机化学学报, 2016, 32(11):2058-2062 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20161122&flag=1CHANG Hui-Qin, YUAN Zhi-Ze, LAI Xiao -Qing, et al. Chinese J. Inorg. Chem., 2016, 32(11):2058-2062 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20161122&flag=1
-
[9]
Liu H, Dong Y, Zhang B, et al. Sens. Actuators B, 2016, 234:616-624 doi: 10.1016/j.snb.2016.04.175
-
[10]
Ponnuvel K, Kumar M, Padmini V. Sens. Actuators B, 2016, 227:242-247 doi: 10.1016/j.snb.2015.12.017
-
[11]
Wu W N, Mao P D, Wang Y, et al. Spectrochim. Acta A, 2018, 188:324-331 doi: 10.1016/j.saa.2017.07.020
-
[12]
Sheldrick G M. SADABS, University of Göttingen, Germany, 1996.
-
[13]
Sheldrick G M. SHELX-97, Program for the Solution and the Refinement of Crystal Structures, University of Göttingen, Germany, 1997.
-
[14]
Huang Y Q, Zhao W, Chen J G, et al. Z. Anorg. Allg. Chem., 2012, 638:679-682 doi: 10.1002/zaac.v638.3/4
-
[15]
Huang Y Q, Wan Y, Chen H Y, et al. New J. Chem., 2016, 40:7587-7595 doi: 10.1039/C6NJ01231K
-
[1]
-
Figure 1 Diamond drawing of 1~3 (a~c) with 30% thermal ellipsoids; Extended 2D supramolecular structure in complex 1 (d); Chain-like structures in complex 2 (e, along c axis) and 3 (f) formed by hydrogen bonds (shown in dashed line), respectively
(a~c) H atoms are omitted for clarity; Symmetry codes: ⅰ x, -1+y, z; ⅱ 1-x, 1-y, -z; ⅲ 1.5-x, -0.5+y, 0.5-z; ⅳ x, 0.5-y, 0.5+z; ⅴ -0.5+x, 1.5-y, -0.5+z
Table 1. Crystal data and structure refinement for complexes 1~3
1 2 3 Empirical formula C12H15N3O7SZn C12H11N3OCl2Cd C12H11N3OI2Cd Formula weight 410.70 396.54 579.44 Size / mm 0.15×0.12×0.06 0.20×0.18×0.16 0.18×0.16×0.15 Crystal system Monoclinic Monoclinic Monoclinic Space group P21/n P21/c P21/n a / nm 1.033 9(7) 0.792 90(19) 0.746 79(12) b / nm 0.664 5(4) 1.523 4(3) 1.659 7(2) c / nm 2.179 5(14) 1.404 0(3) 1.263 00(12) β/(°) 93.915(11) 123.322(10) 90.132 0(10) V/ nm3 1.493 8(16) 1.417 1(5) 1.565 4(3) Z 4 4 4 Dc / (g·cm-3) 1.826 1.859 2.459 Unique 2 616 2 495 2 756 Rint 0.072 7 0.021 1 0.022 0 GOF 1.032 1.070 1.053 R indices [I/>2σ(I)] R1=0.044 3, wR2=0.079 5 R1=0.026 0, wR2=0.063 5 R1=0.026 9, wR2=0.059 7 R indices (all data) R1=0.091 6, wR2=0.092 7 R1=0.031 4, wR2=0.066 8 R1=0.032 7, wR2=0.062 5 Table 2. Selected bond lengths (nm) and angles (°) in complexes 1~3
1 Zn1-N1 0.209 9(4) Zn1-N2 0.205 9(4) Zn1-O1 0.2l5 8(3) Zn1-O2 0.211 4(3) Zn1-O6 0.200 0(3) Zn1-O5ⅰ 0.227 9(3) O6-Zn1-N2 161.91(14) N1 -Zn1 -O2 87.73(14) O6-Zn1-O5ⅰ 86.08(12) O6-Zn1-N1 107.02(15) O6-Zn1 -O1 87.47(14) N2-Zn1-O5ⅰ 86.09(13) N2-Zn1-N1 88.97(15) N2-Zn1 -O1 76.09(15) N1-Zn1-O5ⅰ 87.98(13) O6-Zn1 -O2 95.30(13) N1 -Zn1 -O1 164.90(13) 02-Zn1-O5ⅰ 175.71(12) N2-Zn1 -O2 93.70(13) 02-Zn1 -O1 95.28(13) H-Zn1-O5ⅰ 88.83(12) 2 Cd1-N1 0.235 6(2) Cd1-N2 0.231 4(2) Cd1-O1 0.236 0(2) Cd1-Cl1 0.241 82(11) Cd1-Cl2 0.244 57(10) Cl1-Cd1-Cl2 116.27(4) N2-Cd1-N1 79.00(8) N2-Cd1-Cl1 117.11(7) N2-Cd1-Cl2 126.40(7) N2-Cd1-O1 69.66(8) N1-Cd1-Cl1 97.34(7) N1-Cd1-Cl2 98.92(7) N1-Cd1-O1 147.27(8) O1-Cd1-Cl1 105.00(6) O1-Cd1-Cl2 92.37(6) 3 Cd1-N1 0.237 8(4) Cd1-N2 0.230 8(3) Cd1-O1 0.238 4(3) Cd1-I1 0.274 72(7) Cd1-I2 0.273 21(5) I2-Cd1-I1 109.208(16) N2-Cd1-N1 78.88(12) N2-Cd1-I2 127.43(9) N2-Cd1-I1 122.37(9) N2-Cd1-O1 69.29(12) N1-Cd1-I2 100.14(9) N1-Cd1-I1 102.92(9) N1-Cd1-O1 148.15(12) O1-Cd1-I2 98.28(9) O1-Cd1-I1 95.17(10) Symmetry codes: ⅰ x, -1+y, z Table 3. Hydrogen bonds information for complexes 1~3
D-H …A d(D-H) / nm d(H …A) / nm d(D …A) / nm ∠DHA/(°) 1 O6-H6A …O4ⅰ 0.085 0.191 0.270 9(5) 155.1 N3-H3 …O3ⅱ O.O86 0.193 0.274 4(5) 158.2 O6-H6B …O4ⅲ 0.085 0.189 0.270 1(5) 159.6 O7-H7A …O4 0.085 0.26 0.317 4(6) 125.9 O7-H7B …O3 0.085 0.25 0.297 4(7) 116 2 N3-H3A …Cl2iv 0.086 0.235 0.318 9(3) 164.3 3 N3-H3A …I1v 0.086 0.311 0.383 1(4) 142.3 Symmetry codes: ⅰ x, -1+y, z; ⅱ 1-x, 1-y, -z; ⅲ 1.5-x, -0.5+y, 0.5-z; ⅳ x, 0.5-y, 0.5+z; ⅴ -0.5+x, 1.5-y, -0.5+z -

 扫一扫看文章
扫一扫看文章
计量
- PDF下载量: 3
- 文章访问数: 1342
- HTML全文浏览量: 160


 下载:
下载:



 下载:
下载:

