
一个新颖南极微生物酯酶EST112-2的功能鉴定和在手性叔醇(S)-芳樟醇制备中的应用
English
Functional Characterization of a New Antarctic Microbial Esterase EST112-2 and Its Use in the Preparation of Chiral Tertiary Alcohol (S)-Linalool
-
Chiral tertiary alcohols (TAs) are recognized as key building blocks for the synthesis of a great variety of valuable flavor compounds and pharmaceuticals.[1] For example, chiral linalool is an important chiral tertiary alcohol used in flavor industry and its enantiomers have been found to differ in odor. The (R)-enantiomer of linalool is a major constituent of Cinnamonium camphora and Cayenne linaloe, whereas the (S)-enantiomer of linalool is a major constituent of coriander oil.[2] More importantly, chiral TAs have also been utilized as intermediates for the synthesis of many valuable drugs, such as efavirenz, one drug clinically used for treatment of human immunodeficiency virus (HIV), [3] and taxol, one well-known drug for the treatment of cancers.[4] Chiral linalool has also been proven to exhibit antibacterial and antiviral activities.[5] So, sustainable strategies for the manufacture of optically pure TAs are highly desirable.
Chiral TAs can be synthesized through traditional organic synthesis. However, traditional organic synthesis generally requires highly toxic heavy metal catalysts and suffers from harsh working conditions and great pollutions to the environments.[6] Another route for the synthesis of chiral compounds is through biocatalytic methods, which generally behave much better enantioselectivity than traditional organic synthesis and can work under mild conditions.[7] For example, chiral secondary alcohols can be prepared through the bio-reductions of keto precursors using dehydrogenases as the biocatalysts. While chiral TAs can not be synthesized through bio-reductions using dehydrogenases, due to the lack of corresponding keto precursors.
Another feasible biocatalytic method for the production of TAs is through the kinetic resolutions of racemic precursors using hydrolase such as esterases and lipases.[8] Although enzymatic kinetic resolution, either through trans-esterification reactions or through hydrolysis reactions, has been proven to be an efficient way for the preparation of secondary alcohols, commercially available esterases and lipases generally exhibited low activity and enantioselctivity during kinetic resolutions, possibly because of the steric hindrance caused by the chemical structures of TAs.[9] Thus, it should be highly desirable to identify novel esterases for the preparation of chiral TAs. Oceans are of diverse extreme environments and contain millions of novel microorganisms. It should be highly desirable to identify novel enzymes, such as novel esterases, from marine microorganisms and utilize those novel enzymes in the enzymatic preparation of chiral TAs.
Herein, we identified a new esterase EST112-2 from Psychrobacter sp. HU112 isolated from the antarctic sediments. Esterase EST112-2 was found to contain one conserved GGGX-motif located in the oxyanion binding pocket, which was essential for the kinetic resolution of TAs.[8b] Esterase EST112-2 was further utilized as a green biocatalyst in the preparation of (S)-linalool through asymmetric hydrolysis of racemic linalyl acetate. The enantioselectivity of EST112-2 was much better than those of previous reports about the preparation of chiral linalool through enzymatic kinetic resolutions (Eq. 1).

(1) 1. Results and discussion
1.1 Sequence analysis
Through bio-informatic screening, a 1401-bp gene coding for a putative esterase EST112-2 of 466 amino acid residues was identified from the genomic sequence of Psychrobacter sp. HU112. EST112-2 shared its maximal identity (73%) to one hypothetical protein (WP_055125642) from P. glacincola, one putative esterase (GAF59704) from Psychrobacter sp., and one putative esterase (WP_060492094) from Psychrobacter sp. No signal peptide sequence was found upstream of EST112-2 through SignaIP 4.0 Server, suggesting that EST112-2 was probably an intracellular esterase. A GDSAG motif was identified in the protein sequence of EST112-2. A phylogenetic tree was constructed using the protein sequences of EST112-2 and other lipolytic enzymes, and showed that EST112-2 should be grouped into GDSAG subfamily.[10] Multiple sequence alignment of EST112-2 and other GDSAG subfamily lipolytic enzymes was built. A catalytic triad formed by Ser282-Asp397-H472 and a vital motif GGGX correlated to the hydrolysis of esters of tertiary alcohols was found in the protein sequence of EST112-2.
1.2 Expression and purification of recombinant esterase EST112-2
The recombinant EST112-2 was successfully expressed in E. coli BL21(DE3). 5.2 mg purified recombinant EST112-2 was obtained from 1 L of culture and contained a total activity of 1076 U. No obvious difference of protein expression was observed when induced at 16 ℃ or 22 ℃. A single protein band consistent with its calculated molecular weight (55) could be observed on the SDS-PAGE gel.
1.3 Biochemical characterization of EST112-2
1.3.1 Substrate specificity
p-Nitrophenyl esters with various acyl chain lengths (C2—C14) were used to test the substrate specificity of EST112-2. EST112-2 preferred short-chain p-nitrophenyl esters as the substrate and the highest activity was obtained in the hydrolysis of p-nitrophenyl decanoate (C6). EST112-2 exhibited no hydrolytic activity towards olive oil. Since EST112-2 could hydrolyze short-chain p-nitrophenyl ester instead of water-insoluble long chain fats, EST112-2 was characterized to be an esterase. [11]
1.3.2 Effect of pH on the hydrolytic activity and stability of EST112-2
EST112-2 exhibited its highest hydrolytic activity in 50 mmol/L Tris-HCl buffer at pH 7.5 and exhibited relatively lower hydrolytic activities when the pH was lower than 7.0 or higher than 8.0. EST112-2 was found to be quite stable at pH from 5 to 9.0, with all the residual activities being over 58% of its original activity. EST112-2 was found to be mostly stable at pH=7.5.
1.3.3 Effect of temperature on the hydrolytic activity and stability of EST112-2-2
EST112-2 exhibited relative high activities within a narrow temperature range. EST112-2 exhibited its maximal hydrolytic activity at 40 ℃, but exhibited only 7.6% of its original activity at 50 ℃. The thermo-stability of EST112-2 was studied by pre-incubation at temperatures ranging from 4 ℃ to 60 ℃. The thermo-stability of EST112-2 was also not quite satisfying because EST112-2 rapidly lost its activity at over 50 ℃ and was completely deactivated after incubation at 60 ℃ for 30 min.
1.3.4 Effect of metal ions on the stability of EST112-2
Among the nine metal ions tested, Ca2+ and Mg2+ almost had no influence on the activity of EST112-2, Co2+, Ba2+ and Mn2+ slightly inhibited the activity of EST112- 2, and Cu2+, Zn2+, Ni2+ and Fe2+ severely inhibited the activity of EST112-2.
1.3.5 Effect of detergents on the stability of EST112-2
Non-ionic detergents such as Triton-100, Tween-20 and Tween-80 inhibited the hydrolytic activity of EST112-2. Ionic detergents including SDS and guanidinium chloride also strongly inhibited the hydrolytic activity of EST112-2. EDTA was found to slightly inhibit the hydrolytic activity of EST112-2.
1.3.6 Effect of organic solvents on the stability of EST112-2
As shown in Supplementary Table 2, of all the tested organic solvents, n-hexane, cyclohexane, methanol and dimethyl sulfoxide (DMSO) stimulated the hydrolytic activity of EST112-2. While all the other organic solvents, especially propanol, pentanol, dioxane and benzene had negative effects on the hydrolytic activity of EST112-2.
Table 2
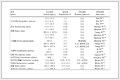 Table 2. Effects of co-solvents on the enantioselective hydrolysis of (±)-linalyl acetate by EST112-2
Table 2. Effects of co-solvents on the enantioselective hydrolysis of (±)-linalyl acetate by EST112-2Organic Co-solvents Concentration (V/V)/% ee/% Yield/% Control 56 80 2.5 53 72 5 68 50 Dichloromethane 10 68 41 15 65 33 20 - - DMSO 10 53 57 n-Hexane 10 47 75 Acetonitrile 10 55 58 Cyclohexane 10 31 78 Ethyl acetate 10 35 59 Methanol 10 50 57 Tetrahydrofuran 10 50 68 1, 4-Dioxane 10 - - Toluene 10 - - Xylene 10 57.5 55 1.4 Kinetic analysis of EST112-2
The kinetic parameters of esterase EST112-2 was calculated with substrate concentration of 0.01~2.00 mmol/L. The specific activity of purified DAEst6 was (207±6) U/mg using p-NP C6 as the substrate, with Km being (2.76±0.08) mmol/L, Vmax being (2.02±0.02) mmol/ (min•mg) and kcat being (189.8±0.6) s-1, respectively.
1.5 Optimization of the kinetic resolution of (±)-linalyl acetate by EST112-2
1.5.1 Effect of temperature on the kinetic resolution of (±)-linalyl acetate by EST112-2
We first investigated the effect of temperature on the kinetic resolution of (±)-linalyl acetate by EST112-2. As shown in Figure 1, the best enantiomeric excess (ee) and yield of product (S)-linalool could reach 53% and 79%, respectively at 30 ℃. The yield sharply decreased as the reaction temperature exceeded 40 ℃. Thus, the optimal temperature for the enantioselective hydrolysis of (±)-linalyl acetate was characterized to be 30 ℃.
Figure 1
1.5.2 Effect of pH on the kinetic resolution of (±)-linalyl acetate by EST112-2
The variation of pH may alter the ionic state of both the substrate and the stereochemical configuration of enzymes, which in turn influences the enzymatic activity and enantioselectivity. As shown in Table 1, the best ee (55%) and yield (80%) could be observed when the kinetic resolution of (±)-linalyl acetate was carried out under in 50 mmol/L Tris-HCl buffer at pH=7.5.
Table 1
pH ee/% Yield/% 5.0a 37 43 6.0a 38 60 6.5b 43 75 7.0b 53 56 7.5b 52 78 7.5c 55 80 8.0b 49 74 8.0c 52 78 8.5c 50 76 9.0c 48 65 a Buffer for 50 mmol/L HAc-NaAc. b Buffer for 50 mmol/L Na2HPO4- NaH2PO4. c Buffer for 50 mmol/L Tris-HCl. 1.5.3 Effect of ionic strength on the kinetic resolution of (±)-linalyl acetate by EST112-2
Ionic strength may also greatly affect the enantioselectivity during enzymatic kinetic resolutions. As shown in Figure 2, the ee and yield remained relative high under buffer concentrations of 0.05~0.6 mol/L, but decreased when the buffer concentrations exceeded 0.6 mol/L. The optimal ionic strength for the best ee (56%) and yield (86%) were found to be 0.1 mol/L.
Figure 2
1.5.4 Effect of co-solvents on the kinetic resolution of (±)-linalyl acetate by EST112-2
The kinetic resolution of (±)-linalyl acetate by EST112- 2 was carried out in various co-solvents at a concentration of 10% (V/V). The highest ee (68%) was obtained when dichloromethane was used as co-solvent at a concentration of 5% (V/V). But the corresponding yield decreased compared with control experiments (Table 2).
1.5.5 Effect of catalyst loading on the kinetic resolution of (±)-linalyl acetate by EST112-2
As shown in Table 3, with the increase of catalyst loading from 20 mg/mL to 60 mg/mL, the yield increased from 62% to 99%, but the ee decreased from 59% to 54%. The increase of the catalyst loading to 80 mg/mL did not shorten the reaction time or improve the enantioselectivity of EST112-2. Thus, 60 mg/mL was chosen as optimal catalyst loading for the kinetic resolution of racemic linaly acetate by EST112-2.
Table 3
Enzyme loading/(mg•mL-1) ee/% Yield/% Reaction time/h 20 59 62 14 40 54 98 14 60 54 99 10 80 51 99 10 1.5.6 Effect of substrate concentration on the kinetic resolution of (±)-linalyl acetate by EST112-2
To test the effect of substrate concentration on the kinetic resolution of (±)-linalyl acetate by EST112-2, (±)-linalyl acetate of different concentrations was added to the enzymatic reactions. As shown in Figure 3, the optimal substrate concentration was found to be 30 mmol/L for the kinetic resolution of (±)-linalyl acetate, with the ee being 54.5% and yield being 99%.
Figure 3
1.5.7 Effect of reaction time on the kinetic resolution of (±)-linalyl acetate by EST112-2
As shown in Figure 4, the highest ee (71%) was obtained at 2 h, but the yield was simply 22%. With the increase of reaction time, the optical purity of product (S)-linalool decreased. When the reaction time reached 6 h, both the ee value (66%) and yield (72%) were acceptable. So the optimal reaction time for the kinetic resolution of (±)-linalyl acetate was found to be 6 h.
Figure 4
1.6 Comparison of the enantioselectivity of EST112-2 with that of other lipases/eaterases in the resolution of (±)-linalyl acetate
Before our study, there were also some reports about the preparation of chiral linalool through the asymmetric hydrolysis of (±)-linalyl acetate by lipases and esterases. As shown in Table 4, lipases/esterases from Bacillus sp. BP-7[12], Rhodococcus sp. CR-53[13], Rhodococcus ruber SM[14], Rhodococcus ruber DSM 43338[14], and Nocardia sp. H8[15] were used to resolve (±)-linalyl acetate and generated chiral linalool. However, in almost all those cases, the enantioselectivities of those esterases/lipases were not high. Thus, it is desirable to identify and utilize novel bioctatalysts for the synthesis of tertiary alcohols with better optical purity. In our work, from the genome of Psychrobacter isolated from the antarctic sediments we identified a new esterase, EST112-2, which exhibts relatively low sequence identifies with the esterases known so far. Through the comparation of EST112-2 with other esterases/lipases in the kinetic resolution of (±)-linalyl acetate, the enantio-preference of EST112-2 was consistence with other reported esterases/lipases and generated (S)-linalool. But both the enantiomeric excess (over 66%) and the yield (over 72%) of product (S)-linalool catalysed by esterase EST112-2 were much higher than previous reports.
Table 4
 Table 4. Comparison of the stereo-selectivity of EST112-2 with other lipases/eaterases in the resolution of (±)-linalyl acetatea
Table 4. Comparison of the stereo-selectivity of EST112-2 with other lipases/eaterases in the resolution of (±)-linalyl acetateaOrigin of enzymes ee/% Yield/
%Time Config. Ref. Bacillus sp BP-7 20 64 0.25 NR [12] Rhodococcus sp. CR-53 39 35 2 NR [13] Rhodococcus ruber SM 58 30 1 S-(+) [14] Nocardia sp. H8 15 31 1 S-(+) [15] Rhodococcus ruber DSM 43338 48 10 16 S-(+) [14] Psychrobacter sp. HU112 66 72 6 S-(+) This study a NR denotes not reported. 2. Conclusions
TAs are crucial building blocks for the synthesis of a great variety of valuable flavors and pharmaceuticals. However, the preparations of chiral tertiary alcohols, especially those with high eanantomeric excess, were notoriously hard. The two enantiomers of linalool, a tertiary alcohol, have been found to differ in odor. The enantiomeric excess and the yield of chiral linalool generated from previous reports were not satisfying, possibly because of the steric hindrance caused by the chemical structure of linalool. We identified and functionally characterized a new esterase EST12-2 from the genome of Psychrobacter sp. HU112 isolated from the antarctic sediments. Esterase EST112-2 was further utilized in the kinetic resolution of (±)-linalyl acetate and generated (S)-linalool with much higher enantiomeric excess of over 66% and a yield of over 72%. In conclusion, Antarctic microbial esterase EST112-2 is a promising green biocatalyst in the preparation of chiral tertiary alcohols represented by linalool. Next, we will try to further imporve the enanio-selectivity and production yield of EST112-2 through enzyme engineering and also try to identify new esterases for the preparation of chiral tertiary alcohols through kinetic resolution.
3. Experimental section
3.1 Strains, vectors and chemicals
The strain Psychrobacter sp. HU112 was isolated from the Antarctic sediments. Psychrobacter sp. HU112 was grown in the high-salt Luria-Bertani medium (HLB) (peptone 1%, yeast extract 0.5%, NaCl 2%) at 28 ℃ for overnight. The plasmid pET-28a (+) (Novagen, USA) was used as the expression vector. The strains Escherichia coli DH5α and E. coli BL21 (DE3) were used for routine cloning and gene expression, respectively. DNA purification kits and FastDigest restriction endonucleases were purchased from Thermo Fisher Scientific. T4 DNA ligase and FastPfu polymerases were purchased from Transgene (Beijing, China). Substrates p-nitrophenyl acetate (p-NPA, C2), p-nitrophenyl butyrate (p-NPB, C4), p-nitrophenyl decanoate (p-NPD, C6), p-nitrophenyl caprylate (p-NPC, C8), p-nitrophenyl laulate (p-NPL, C12), and p-nitrophenyl palmitate (p-NPP, C16) were obtained from Sigma-Aldrich (St Louis, MO, USA). (±)-linalool, (R)-(-)-linalool and (±)-linalyl acetate were purchased from Aladdin Chemistry Corporation.
3.2 Gene cloning and sequence analysis
The genome sequence of Psychrobacter sp. HU112 was completed by Majorbio LLC, Shanghai. Based on the coding DNA sequence of a putative esterase (here labeled as EST112-2) from the genome of Psychrobacter sp. HU112, two degenerate primers (F:5'-CACGGATCCATGTCT- CACTCAACCATATCGCTCAATACC-3' and R:5'-CC- GCTCGAGTTAGGCCGCTAACTGCCC-3') were designed (restriction sites for BamH I and Xho I were underlined). PCR product was digested by BamH I and Xho I according to the manufacturers instruction, purified using DNA purification kits, and cloned into vector pET28a(+). The expression plasmid pET28a-EST112-2 was confirmed by sequencing from Majorbio Ltd (Shanghai, China).
Multiple sequence alignments were constructed using the DNAMAN 7.0 program. Three-dimensional model of enzymes was built and analyzed through the online SwissModle server at https://swissmodel.expasy.org. Phylogenetic analysis and molecular evolutionary analysis were conducted using MEGA software version 5. Secretion signal was predicted by the SignalP 4.0 prediction tool from http://www.cbs.dtu.dk/services/SignalP/. The theoretical molecular weight and pI of proteins were estimated by expasy from http://web.expasy.org/compute_pi/.
3.3 Expression and purification of recombinant esterase EST112-2
Esterase EST112-2 were heterologous express in E. coli BL21 (DE3). The recombinant cells were cultivated in Luria-Bertani medium harboring 50 μg/mL of kanamycin at 37 ℃ until the OD600 reached 0.6. Then, the cells were induced at 16 ℃ or 22 ℃ with 0.2 mmol/L IPTG. To purify EST112-2, 1 L of LB medium culture was harvested by centrifugation at 4000 r/min for 20 min at 4 ℃. The cells were dissolved in 50 mmol/L Tris-HCl buffer (pH 8.0), and disrupted by sonication on ice. The supernatants were collected by centrifugation at 12000 r/min for 20 min. Recombinant EST112-2 was purified using Ni-NTA resins, then imidazole and ions were removed by GE PD-10 desalting column (GE Healthcare Life Sciences). Samples were collected for SDS-PAGE analysis.
3.4 Biochemical characterization of EST112-2
3.4.1 Substrate specificity
The substrate specificity of EST112-2 was measured using 10 mmol/L p-NP esters with acyl chains of different lengths (C2~C16) as the substrates.[16] Standard enzymatic reactions were carried out as mentioned above and were stopped by adding 200-μL n-propanol and measured immediately at 405 nm.
3.4.2 Effect of pH on the activity and stability of EST112-2-2
The optimum pH conditions were determined at 30 ℃ in 50 mmol/L buffers under standard assay conditions: acetic acid/sodium acetate (pH=5.0~6.0), potassium phosphate (pH=6.5~8.0), Tris-HCl (pH=7.5~9.0) and glycine/ NaOH (pH=9.0~10.0). The reactions were stopped by adding 200-μL n-propanol and measured immediately at 405 nm.
To determine the pH stability of EST112-2, 500 U of purified EST112-2 was incubated for 12 h at 4 ℃ in 940-μL 50 mmol/L buffers of different pH. Then 10-μL of 10 mmol/L p-NPD, 40-μL of ethanol incubated at 30 ℃ for 3 min. The enzymatic reactions were stopped by adding 200-μL n-propanol and measured immediately at 405 nm.
3.4.3 Effect of temperature on the activity and stability of EST112-2
The optimal temperature of EST112-2 was determined by measuring the esterase activity at different temperatures (4~60 ℃) in 50 mmol/L Tris-HCl buffer (pH 7.5) under standard assay conditions. To investigate the thermo-stability of EST112-2 (500 U) of purified EST112-2 was incubated at different temperatures (4~60 ℃) for 1 h. Then the residual activities of EST112-2 were measured under standard assay conditions as described previously. The enzymatic reactions with untreated EST112-2 were utilized as control experiments.
3.4.4 Effect of metal ions on the stability of EST112-2
The effect of various metal ions on the esterase activity was assessed by incubating EST112-2 (500U) in 50 mmol/L Tris-HCl, pH=7.5, and 5 mmol/L of different metal ions at 4 ℃ for 12 h. Then, the residual activities of EST112-2 were measured under standard assay conditions as previously described. The enzymatic reactions with untreated EST112-2 were utilized as control experiments.
3.4.5 Effect of detergents on the stability of EST112-2
For the determination of detergents on the stability of EST112-2 (500 U) purified esterase was incubated in the presence of various detergents: Tween-20, Tween-80 and Triton-100 at a final concentration of 0.025% (V/V); Guanidinium chloride, Urea, SDS and EDTA at final concentrations of 2 and 5 mmol/L for 12 h at 4 ℃, respectively. Then, the residual activities of EST112-2 were measured under standard assay conditions as described previously. Enzymatic reactions with untreated EST112-2 were utilized as control experiments.
3.4.6 Effect of organic solvents on the stability of EST112-2
To investigate the effect of organic solvents on the stability of EST112-2, 500 U purified esterase was pre-incubated with different organic solvents at a final concentration of 20% (V/V) for 12 h at 35 ℃. The residual activities of EST112-2 were measured under standard assay conditions as previously described. The enzymatic reactions with untreated EST112-2 were utilized as control experiments.
3.5 Kinetic analysis of EST112-2
The kinetic parameters of EST112-2 were calculated at 35 ℃, pH 7.5 and 50 mmol/L Tris-HCl buffer containing p-NPD of concentrations ranging from 0.01 mmol/L to 2 mmol/L. Km and Vmax were obtained using a Lineweaver-Burk plot under the optimal reaction conditions.
3.6 Optimization of kinetic resolution of (±)-linalyl acetate by EST112-2
3.6.1 Effect of temperature on the kinetic resolution of (±)-linalyl acetate by EST112-2
A standard reaction system contained EST112-2 enzyme powder (10 mg), 498-μL Tris-HCl buffer (50 mmol/L, pH=7.5), 20 mmol/L (±)-linalyl acetate. The effect of temperature on the kinetic resolution of (±)-linalyl acetate was performed at 4~60 ℃ for 12 h. After the termination of enzymatic reactions, 500-μL ethyl acetate was added to extract both the substrate and the products. Linalyl acetate and linalool was further analyzed by chiral GC. The enantiomeric excess (ee) of (S)-linalool and yield of (S)-linalool (Y) were calculated by using the equation of Chen et al.[17]
3.6.2 Effect of pH on the kinetic resolution of (±)- linalyl acetateby EST112-2
A standard 500-μL reaction system contained EST112-2 enzymatic powder (10 mg), 20 mmol/L (±)-linalyl acetate and 498-μL buffer with pH ranging from 5.0 to 10.0. After incubating the enzymatic reactions for 12 h at 30 ℃, 500-μL ethyl acetate was added to extract the substrate and products. Linalyl acetate and linalool were further analyzed by chiral GC.
3.6.3 Effect of ionic strength on the kinetic resolution of (±)-linalyl acetate by EST112-2
The effect of ionic strength (0.05~1 mol/L) on the kinetic resolution of racemic linalyl acetate was investigated using a 500-μL enzymatic reactions harboring EST112-2 enzyme powder (10 mg), 498-μL Tris-HCl buffer (100 mmol/L, pH=7.5), and 20 mmol/L (±)-linalyl acetate. After incubating the enzymatic reactions for 12 h at 30 ℃, 500-μL ethyl acetate was added for extraction. Both linalyl acetate and linalool were further analyzed by chiral GC.
3.6.4 Effect of co-solvents on the kinetic resolution of (±)-linalyl acetate by EST112-2
The effect of different co-solvents on the kinetic resolution of (±)-linalyl acetate by EST112-2 was analyzed using a reaction system containing EST112-2 enzymatic powder (10 mg), 448-μL Tris-HCl buffer (100 mmol/L, pH=7.5), 20 mmol/L (±)-linalyl acetate and 10% (V/V) different organic solvents. Afterwards, the concentrations of co-solvents ranging from 5% to 20% (V/V) were optimized. After incubating the enzymatic reactions for 12 h at 30 ℃, 500-μL ethyl acetate was added into enzymatic reactions for extraction. Both linalyl acetate and linalool were further analyzed by chiral GC.
3.6.5 Effect of catalyst loading on the kinetic resolution of (±)-linaly acetate by EST112-2
The effect of catalyst loading on the kinetic resolution of (±)-linaly acetate by EST112-2 was analyzed by a reaction system, contained EST112-2 enzymatic powder (10~40 mg), 448 μL of Tris-HCl buffer (100 mmol/L, pH=7.5), 20 mmol/L (±)-linalyl acetate and 10% (V/V) CH2Cl2. After incubating the enzymatic reactions for 12 h at 30 ℃, 500-μL ethyl acetate was added for extraction. Both linalyl acetate and linalool were further analyzed by chiral GC.
3.6.6 Effect of substrate concentration on the kinetic resolution of (±)-linalyl acetate by EST112-2
The effect of substrate concentration on the kinetic resolution of (±)-linalyl acetate catalyzed by EST112-2 was analyzed using a reaction system containing EST112-2 enzymatic powder (10 mg), 448-μL Tris-HCl buffer (100 mmol/L, pH 7.5), 10% (V/V) CH2Cl2 and (±)-linaly acetate (5~50 mmol/L). After incubating the enzymatic reactions for 10 h at 30 ℃, 500-μL ethyl acetate was added for extraction. Both linalyl acetate and linalool was further analyzed by chiral GC.
3.6.7 Effect of reaction time on the kinetic resolution of (±)-linalyl acetate by EST112-2
The effect of reaction time on the kinetic resolution of (±)-linalyl acetate catalyzed by EST112-2 was analyzed by using a 500-μL reaction system containing EST112-2 enzymatic powder (30 mg), 100 mmol/L Tris-HCl buffer (pH 7.5), 10% (V/V) CH2Cl2 and 30 mmol/L (±)-linalyl acetate. Linalyl acetate and linalool in the reactions were monitored every 2 h and further analyzed by chiral GC.
3.7 Gas chromatograph analysis
The residual linalyl acetate and generated linalool in enzymatic reaction system was analyzed via gas chromatograph (FULI GC-9790 Ⅱ) using a equipped 112-6632 CYCLOSIL-B chiral capillary column (30 m×0.25 mm ID, 0.25 μm df). The temperatures of H2 flame ionization detector and injector were set at 250 and 280 ℃, respectively. Nitrogen served as carrier gas at a split flow rate of 1.20 mL/min. The oven temperature was held at 100 ℃ for 1 min, then increased at 10 ℃/min to 220 ℃ and held for 3 min.
Supporting Information The protein sequence analysis and biochemical characterization. The Supporting Information is available free of charge via the Internet at http://sioc-journal.cn.
-
-
[1]
(a) Yao, G. ; Haque, S. ; Sha, L. ; Kumaravel, G. ; Wang, J. ; Engber, T. M. ; Whalley, E. T. ; Conlon, P. R. ; Chang, H. ; Kiesman, W. F. ; Petter, R. C. Biorg. Med. Chem. Lett. 2005, 15, 511.
(b) Friel, D. K. ; Snapper, M. L. ; Hoveyda, A. H. J. Am. Chem. Soc. 2008, 130, 9942.
(c) Shibasaki, M. ; Kanai, M. Chem. Rev. 2008, 108, 2853. -
[2]
Frizzo, C. D.; Santos, A. C.; Paroul, N.; Serafini, L. A.; Dellacassa, E.; Lorenzo, D.; Moyna, P. Braz. Arch. Biol. Technol. 2000, 43, 313. doi: 10.1590/S1516-89132000000300011
-
[3]
(a) Thompson, A. S. ; Corley, E. G. ; Huntington, M. F. ; Grabowski, E. J. J. Tetrahedron Lett. 1995, 36: 8937.
(b) Gulick, R. M. ; Ribaudo, H. J. ; Shikuma, C. M. ; Lustgarten, S. ; Squires, K. E. ; Meyer Ⅲ, W. A. ; Acosta, E. P. ; Schackman, B. R. ; Pilcher, C. D. ; Murphy, R. L. ; Maher, W. E. ; Witt, M. D. ; Reichman, R. C. ; Snyder, S. ; Klingman, K. L. ; Kuritzkes, D. R. N. Engl. J. Med. 2004, 350, 1850. -
[4]
Ishihara, Y.; Baran, P. S. Synlett 2010, 41, 1733. https://es.scribd.com/document/19982642/Merck-Index-Name-Reactions
-
[5]
(a) Freires, I. ; Denny, C. ; Benso, B; de Alencar, S. ; Rosalen, P. Molecules 2015, 20, 7329.
(b) Herman, A. ; Tambor, K., Herman, A. Curr. Microbiol. 2016, 72, 165. -
[6]
Noyori, R.; Kitamura, M. Angew. Chem., Int. Ed. 1991, 30, 49. doi: 10.1002/(ISSN)1521-3773
-
[7]
Adrio, J. L.; Demain, A. L. Biomolecules 2014, 4, 117. doi: 10.3390/biom4010117
-
[8]
(a) Holt, J. ; Arends, I. W. C. E. ; Minnaard, A. J. ; Hanefeld, U. Adv. Synth. Catal. 2007, 349, 1341.
(b) Kourist, R. ; Bornscheuer, U. T. Appl. Microbiol. Biotechnol. 2011, 91, 505.
(c) Yan, Q. J. ; Yang, S. Q. ; Duan, X. J. ; Xu, H. B. ; Liu, Y. ; Jiang, Z. Q. J. Mol. Catal. B: Enzym. 2014, 109, 76. -
[9]
Pogorevc, M.; Faber, K. J. Mol. Catal. B:Enzym. 2000, 10, 357. doi: 10.1016/S1381-1177(99)00121-6
-
[10]
(a) Timmis, K. E. Handbook of Hydrocarbon and Lipid Microbiology, Eds. : McGenity, T. J. ; van der Meer, J. R. ; de Lorenzo, V. ; Berlin, Heidelberg, 2010, p. 1099.
(b) Li, P. Y. ; Chen, X. L. ; Ji, P. ; Li, C, Y. ; Wang, P. ; Zhang, Y. ; Xie, B. B. ; Qin, Q. L. ; Su, H. N. ; Zhou, B. C. ; Zhang, Y. Z. ; Zhang, X. Y. J. Biol. Chem. 2015, 290, 11188. -
[11]
(a) Helistö, P. ; Korpela, T. Enzym. Microb. Technol. 1998, 23, 113.
(b) Kulkarni, N. ; Gadre, R. V. J. Ind. Microbiol. Biotechnol. 2002, 28, 344. -
[12]
Fillat, A.; Romea, P.; Urpí, F.; Pastor, F. I. J.; Diaz, P. Appl. Microbiol. Biotechnol. 2014, 98, 4479. doi: 10.1007/s00253-013-5458-9
-
[13]
Bassegoda, A.; Fillat, A.; Pastor, F. I.; Diaz, P. Appl. Microbiol. Biotechnol. 2013, 97, 8559. doi: 10.1007/s00253-012-4676-x
-
[14]
Osprian, I.; Steinreiber, A.; Mischitz, M.; Faber, K. Biotechnol. Lett. 1996, 18, 1331. doi: 10.1007/BF00129965
-
[15]
Pogorevc, M.; Strauss, T. U.; Hayn, M.; Faber, K. Monatsh. Chem. 2000, 131, 639. doi: 10.1007/s007060070092
-
[16]
Winkler, U. K.; Stuckmann, M. J. Bacteriol. 1979, 138, 663.
-
[17]
Chen, C. S.; Fujimoto, Y.; Girdaukas, G.; Sih, C. J. J. Am. Chem. Soc. 1982, 104, 7294. doi: 10.1021/ja00389a064
-
[1]
-
Table 2. Effects of co-solvents on the enantioselective hydrolysis of (±)-linalyl acetate by EST112-2
Organic Co-solvents Concentration (V/V)/% ee/% Yield/% Control 56 80 2.5 53 72 5 68 50 Dichloromethane 10 68 41 15 65 33 20 - - DMSO 10 53 57 n-Hexane 10 47 75 Acetonitrile 10 55 58 Cyclohexane 10 31 78 Ethyl acetate 10 35 59 Methanol 10 50 57 Tetrahydrofuran 10 50 68 1, 4-Dioxane 10 - - Toluene 10 - - Xylene 10 57.5 55 Table 1. Effect of pH on the kinetic resolution of (±)-linalyl acetate by EST112-2
pH ee/% Yield/% 5.0a 37 43 6.0a 38 60 6.5b 43 75 7.0b 53 56 7.5b 52 78 7.5c 55 80 8.0b 49 74 8.0c 52 78 8.5c 50 76 9.0c 48 65 a Buffer for 50 mmol/L HAc-NaAc. b Buffer for 50 mmol/L Na2HPO4- NaH2PO4. c Buffer for 50 mmol/L Tris-HCl. Table 3. Effect of enzyme loading on the enantioselective hydrolysis of (±)-linalyl acetate
Enzyme loading/(mg•mL-1) ee/% Yield/% Reaction time/h 20 59 62 14 40 54 98 14 60 54 99 10 80 51 99 10 Table 4. Comparison of the stereo-selectivity of EST112-2 with other lipases/eaterases in the resolution of (±)-linalyl acetatea
Origin of enzymes ee/% Yield/
%Time Config. Ref. Bacillus sp BP-7 20 64 0.25 NR [12] Rhodococcus sp. CR-53 39 35 2 NR [13] Rhodococcus ruber SM 58 30 1 S-(+) [14] Nocardia sp. H8 15 31 1 S-(+) [15] Rhodococcus ruber DSM 43338 48 10 16 S-(+) [14] Psychrobacter sp. HU112 66 72 6 S-(+) This study a NR denotes not reported. -

 扫一扫看文章
扫一扫看文章
计量
- PDF下载量: 9
- 文章访问数: 1687
- HTML全文浏览量: 214

 下载:
下载:




 下载:
下载:
 下载:
下载:

