Citation:
Qing ZOU, Qiu-Shuang GAO, Peng YAO, Qi-Zheng YAO, Ji ZHANG. Structure-based Screening for the Non-zinc-chelating Selective MMP-13 Inhibitors of Natural Products[J]. Chinese Journal of Structural Chemistry,
;2020, 39(11): 1990-2000.
doi:
10.14102/j.cnki.0254–5861.2011–2831

-
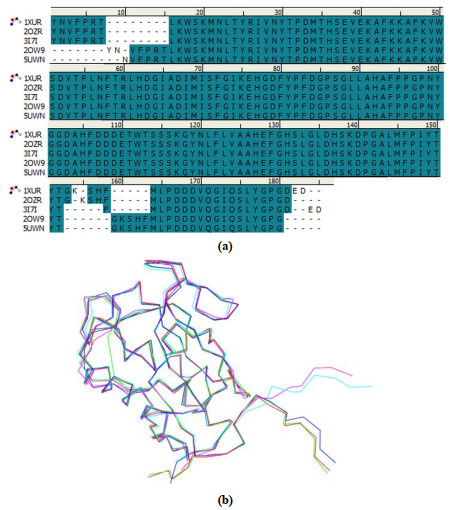
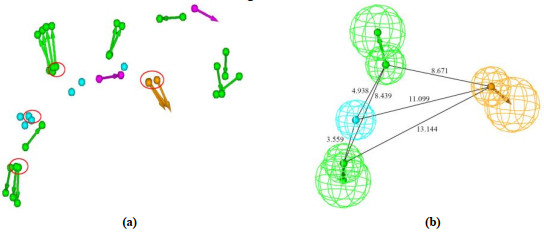
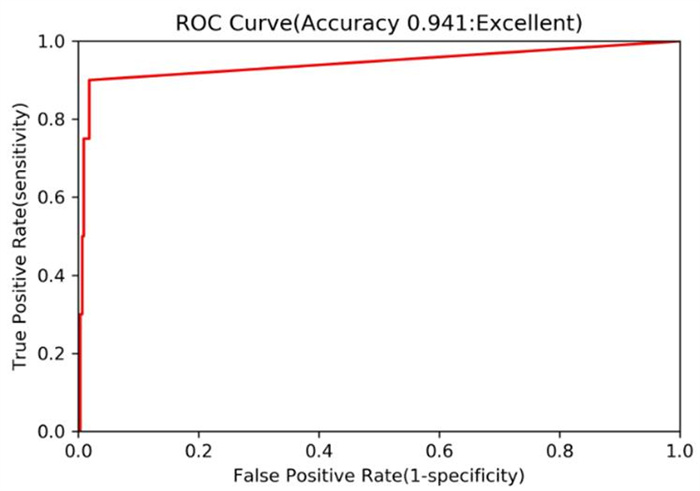
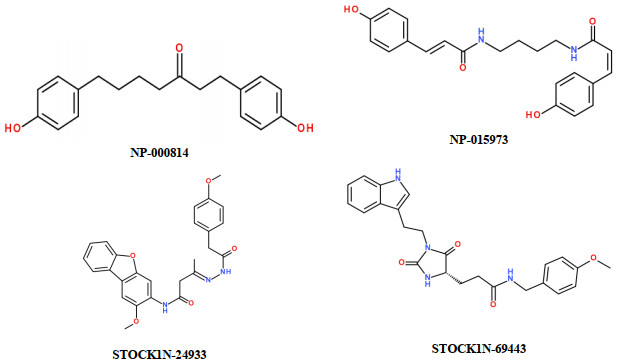
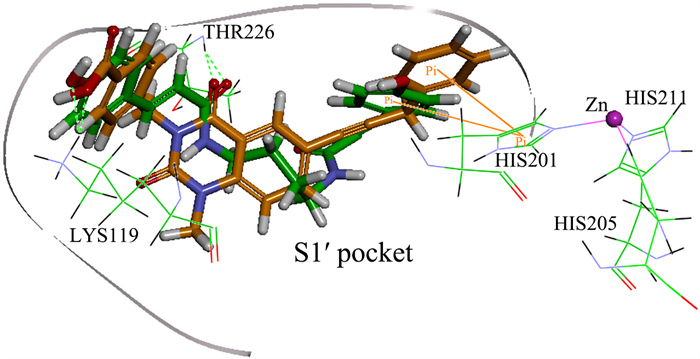
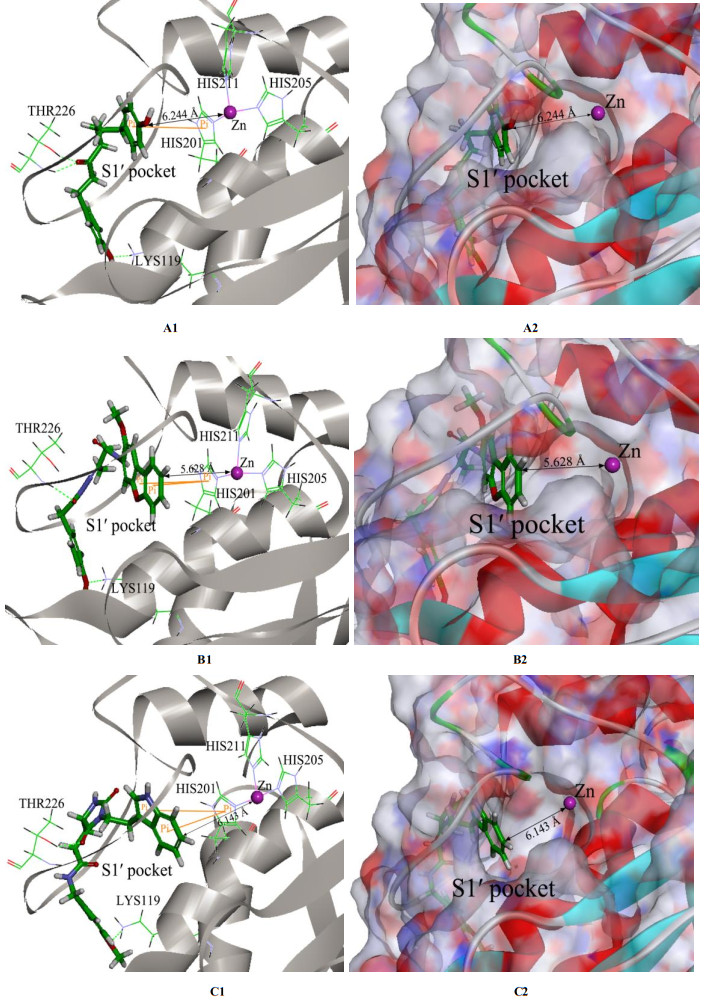
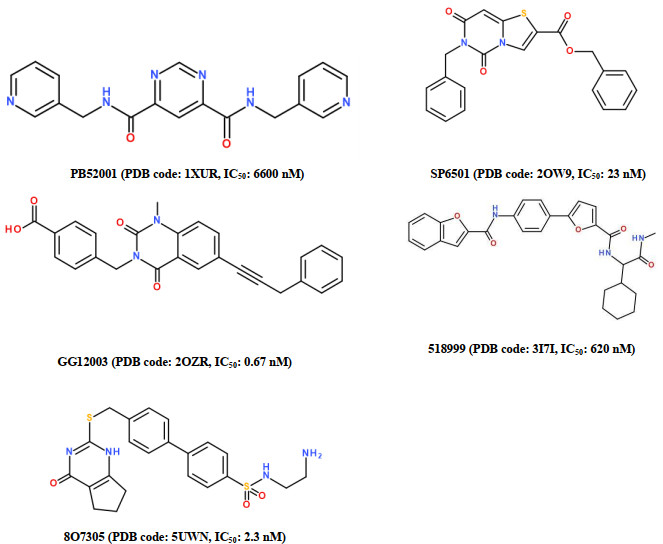
Matrix metalloproteinase-13 (MMP-13) has been considered as a promising therapeutic target for osteoarthritis. In this work, the experimental crystal structures of five MMP-13-ligand complexes are used to build the multiple structure-based pharmacophore model of MMP-13 inhibitors. The reliability of pharmacophore model is validated using a decoy set. The pharmacophore model contains four chemical features: two hydrogen bond acceptor (HBA), one hydrophobic (HY) feature, and one ring aromatic (RA) feature. Particularly, the HY feature is found to orient the MMP-13 inhibitors deep into the S1' pocket of MMP-13 to produce selective inhibition. By carrying out the screening of pharmacophore model and subsequent molecular docking, the four non-zinc-chelating selective MMP-13 inhibitors of natural products (NP-015973, NP-000814, STOCK1N-24933, and STOCK1N-69443) are identified. It is found that the binding modes of MMP-13 with our screened four natural products are very similar to the reported experimental binding mode of MMP-13 with the most active inhibitor (GG12003, IC50: 0.67 nM), and each of them involves the interactions of a ligand with the three amino acid residues Thr226, Lys119, and His201 of MMP-13 receptor. This shows that our modeling results are in good agreement with the relevant experimental results, which strongly supports our screened MMP-13 inhibitors of natural products. These screened natural products may be used as the lead compounds of MMP-13 inhibitors in the future studies of structural modifications.
-

-
-
[1]
Stamenkovic, I. Extracellular matrix remodelling: the role of matrix metalloproteinases. J. Pathol. 2003, 200, 448−464. doi: 10.1002/path.1400
-
[2]
Murphy, G.; Knauper, V.; Atkinson, S.; Butler, G.; English, W.; Hutton, M. Matrix metalloproteinases in arthritic disease. Arthritis Res. 2002, 4, S39−S49. doi: 10.1186/ar572
-
[3]
Conrozier, T.; Ferrand, F.; Poole, A. R.; Verret, C. Differences in biomarkers of type II collagen in atrophic and hypertrophic osteoarthritis of the hip: implications for the differing pathobiologies. Osteoarthr Cartilage. 2007, 15, 462−467. doi: 10.1016/j.joca.2006.09.002
-
[4]
Buckwalter, J. A.; Martin, J. A. Osteoarthritis. Adv. Drug Delivery Rev. 2006, 58, 150−167. doi: 10.1016/j.addr.2006.01.006
-
[5]
Neuhold, L. A.; Killar, L.; Zhao, W.; Sung, M. A.; Warner, L.; Kulik, J.; Turner, J. Postnatal expression in hyaline cartilage of constitutively active human collagenase-3 (MMP-13) induces osteoarthritis in mice. J. Clin. Invest. 2001, 107, 35−44. doi: 10.1172/JCI10564
-
[6]
Stickens, D.; Behonick, D. J.; Ortega, N.; Heyer, B.; Hartenstein, B.; Yu, Y.; Fosang, A. J.; Angel, P.; Werb, Z. Altered endochondral bone development in matrix metalloproteinase 13-deficient mice. Development 2004, 131, 5883−5895. doi: 10.1242/dev.01461
-
[7]
Tallant, C.; Marrero, A. Matrix metalloproteinases: fold and function of their catalytic domains. Biochim. Biophys. Acta 2010, 1803, 20−28. doi: 10.1016/j.bbamcr.2009.04.003
-
[8]
Renkiewicz, R.; Qiu, L.; Lesch, C.; Sun, X.; Devalaraja, R.; Cody, T.; Kaldjian, E.; Welgus, H.; Baragi, V. Broad-spectrum matrix metalloproteinase inhibitor marimastat-induced musculoskeletal side effects in rats. Arthritis Rheum. 2003, 48, 1742−1749. doi: 10.1002/art.11030
-
[9]
Fabre, B.; Ramos, A. Targeting matrix metalloproteinases: exploring the dynamics of the S1' pocket in the design of selective, small molecule inhibitors. J. Med. Chem. 2014, 57, 10205−10219. doi: 10.1021/jm500505f
-
[10]
Engel, C. K.; Pirard, B.; Schimanski, S.; Kirsch, R.; Habermann, J.; Klingler, O.; Schlotte, V.; Weithmann, K. U.; Wendt, K. U. Structural basis for the highly selective inhibition of MMP-13. Chem. Biol. 2014, 12, 181−189.
-
[11]
Johnson, A. R.; Pavlovsky, A. G.; Ortwine, D. F.; Prior, F.; Man, C. F.; Bornemeier, D. A.; Banotai, C. A. Discovery and characterization of a novel inhibitor of matrix metalloprotease-13 that reduces cartilage damage in vivo without joint fibroplasia side effects. J. Biol. Chem. 2007, 282, 27781−27791. doi: 10.1074/jbc.M703286200
-
[12]
Heim-Riether, A.; Taylor, S. J.; Liang, S.; Gao, D. A. Improving potency and selectivity of a new class of non-Zn-chelating MMP-13 inhibitors. Bioorg. Med. Chem. Lett. 2009, 19, 5321−5324. doi: 10.1016/j.bmcl.2009.07.151
-
[13]
Choi, J. Y.; Fuerst, R.; Knapinska, A. M. Structure-based design and synthesis of potent and selective matrix metalloproteinase 13 inhibitors. J. Med. Chem. 2017, 60, 5816−5825. doi: 10.1021/acs.jmedchem.7b00514
-
[14]
Gao, D. A.; Xiong, Z.; Heim-Riether, A.; Amodeo, L.; August, E. M.; Cao, X.; Ciccarelli, L. SAR studies of non-zinc-chelating MMP-13 inhibitors: improving selectivity and metabolic stability. Bioorg. Med. Chem. Lett. 2010, 20, 5039−5043. doi: 10.1016/j.bmcl.2010.07.036
-
[15]
Gege, C.; Bao, B.; Bluhm, H.; Boer, J.; Gallagher, B. M.; Korniski, B.; Powers, T. S.; Steeneck, C. Discovery and evaluation of a non-zn chelating, selective matrix metalloproteinase 13 (MMP-13) inhibitor for potential intra-articular treatment of osteoarthritis. J. Med. Chem. 2012, 55, 709−716. doi: 10.1021/jm201152u
-
[16]
Li, J. J.; Nahra, J.; Johnson, A. R.; Bunker, A.; O'Brien, P.; Yue, W. S.; Ortwine, D. F.; Man, C. F.; Baragi, V. Quinazolinones and pyrido [3, 4-d] pyrimidin-4-ones as orally active and specific matrix metalloproteinase-13 inhibitors for the treatment of osteoarthritis. J. Med. Chem. 2008, 51, 835−841. doi: 10.1021/jm701274v
-
[17]
Taylor, S. J.; Abeywardane, A.; Liang, S.; Muegge, I.; Padyana, A.; Xiong, Z.; Hill-Drzewi, M.; Farmer, B. Fragment based discovery of indole inhibitors of matrix metalloproteinase-13. J. Med. Chem. 2011, 54, 8174−8187. doi: 10.1021/jm201129m
-
[18]
Savi, C. D.; Morley, A. D.; Ting, A.; Nash, I.; Karabelas, K.; Wood, C. M.; James, M.; Norris, S. J.; Karoutchi, G.; Rankine, N. Selective non zinc binding inhibitors of MMP13. Bioorg. Med. Chem. Lett. 2011, 21, 4215−4219. doi: 10.1016/j.bmcl.2011.05.075
-
[19]
Spicer, T. P.; Jiang, J.; Taylor, A. B.; Choi, J. Y.; Hart, P. J.; Roush, W. R.; Field, G. B.; Hodder, P. S.; Minond, D. Characterization of selective exosite-binding inhibitors of matrixmetalloproteinase 13 that prevent articular cartilage degradation in vitro. J. Med. Chem. 2014, 57, 9598−9611. doi: 10.1021/jm501284e
-
[20]
Cragg, G. M.; Grothaus, P. G.; Newman, D. J. New horizons for old drugs and drug leads. J. Nat. Prod. 2014, 77, 703−723. doi: 10.1021/np5000796
-
[21]
Cragg, G. M.; Grothaus, P. G.; Newman, D. Impact of natural products on developing new anti-cancer agents. J. Chem. Rev. 2009, 109, 3012−3043. doi: 10.1021/cr900019j
-
[22]
Yang, S. Y. Pharmacophore modeling and applications in drug discovery: challenges and recent advances. Drug Discovery Today 2010, 15, 444−450. doi: 10.1016/j.drudis.2010.03.013
-
[23]
Gagnon, J. K.; Law, S. M.; Brooks, C. L. Flexible CDOCKER: development and application of a pseudo-explicit structure-based docking method within CHARMM. J. Comput. Chem. 2016, 37, 753−762. doi: 10.1002/jcc.24259
-
[24]
Discovery Studio, Version 3.0; Accelrys Inc: San Diego 2010.
-
[25]
Hamza, A.; Wei, N. N.; Zhan, C. G. Ligand-based virtual screening approach using a new scoring function. J. Chem. Inf. Model 2012, 52, 963−974. doi: 10.1021/ci200617d
-
[26]
Triballeau, N.; Acher, F.; Brabet, I. Virtual screening workflow development guided by the "receiver operating characteristic" curve approach, application to high-throughput docking on metabotropic glutamate receptor subtype 4. J. Med. Chem. 2005, 48, 2534−2547. doi: 10.1021/jm049092j
-
[27]
Hein, M.; Zilian, D. Docking compared to 3D pharmacophores: the scoring function challenge. Drug Discovery Today: Technol. 2011, 7, e229−e236.
-
[28]
Wang, Y. J.; Yang, L. M.; Hou, J. Y.; Zou, Q.; Gao, Q.; Yao, W. H.; Yao, Q. Z.; Zhang, J. Hierarchical virtual screening of the dual MMP-2/HDAC-6 inhibitors from natural products based on pharmacophore models and molecular docking. J. Biomol. Struct. Dyn. 2019, 37, 649−670. doi: 10.1080/07391102.2018.1434833
-
[29]
Hou, J. Y.; Zou, Q.; Wang, Y. J.; Gao, Q.; Yao, W. H.; Yao, Q. Z.; Zhang, J. Screening for the selective inhibitors of MMP-9 from natural products based on pharmacophore modeling and molecular docking in combination with bioassay experiment, hybrid QM/MM calculation, and MD simulation. J. Biomol. Struct. Dyn. 2019, 37, 3135−3149. doi: 10.1080/07391102.2018.1509019
-
[30]
Li, J.; Zhao, F.; Li, M. Z.; Chen, L. X.; Qiu, F. Diarylheptanoids from the Rhizomes of Curcuma kwangsiensis. J. Nat. Prod. 2010, 73, 1667−1671. doi: 10.1021/np100392m
-
[31]
Kim, S. B.; Liu, Q.; Ahn, J. H.; Jo, Y. H.; Turk, A.; Hong, I. P.; Han, S. M.; Hwang, Y. B.; Lee, M. K. Polyamine derivatives from the bee pollen of Quercus mongolica with tyrosinase inhibitory activity. Bioorg. Chem. 2018, 81, 127−133. doi: 10.1016/j.bioorg.2018.08.014
-
[1]
-

-
-
[1]
Jia Chen , Yun Liu , Zerong Long , Yan Li , Hongdeng Qiu . Colorimetric detection of α-glucosidase activity using Ni-CeO2 nanorods and its application to potential natural inhibitor screening. Chinese Chemical Letters, 2024, 35(9): 109463-. doi: 10.1016/j.cclet.2023.109463
-
[2]
Wenchao Chen , Xiaoting Niu , Xinyu Liang , Guanghao Zhu , Xue Zhang , Qing Gong , Cen Xie , Guangbo Ge , Ping Wang , Yuqi He . Rationally engineered a universal far-red fluorogenic substrate of bile salt hydrolase for functional imaging and inhibitor screening. Chinese Chemical Letters, 2026, 37(4): 111320-. doi: 10.1016/j.cclet.2025.111320
-
[3]
Bingwei Wang , Yihong Ding , Xiao Tian . Benchmarking model chemistry composite calculations for vertical ionization potential of molecular systems. Chinese Chemical Letters, 2025, 36(2): 109721-. doi: 10.1016/j.cclet.2024.109721
-
[4]
Haijun Shen , Yi Qiao , Chun Zhang , Yane Ma , Jialing Chen , Yingying Cao , Wenna Zheng . A matrix metalloproteinase-sensitive hydrogel combined with photothermal therapy for transdermal delivery of deferoxamine to accelerate diabetic pressure ulcer healing. Chinese Chemical Letters, 2024, 35(12): 110283-. doi: 10.1016/j.cclet.2024.110283
-
[5]
Chunru Zhao , Yi Liu , Shilong Li , Xiang Wu , Jinghai Liu . PVP decorated H3.78V6O13 microspheres assembled by nanosheets for aqueous zinc ion batteries at variable work temperature. Chinese Chemical Letters, 2025, 36(6): 110185-. doi: 10.1016/j.cclet.2024.110185
-
[6]
Hongmei Yu , Baoxi Zhang , Meiju Liu , Cheng Xing , Guorong He , Li Zhang , Ningbo Gong , Yang Lu , Guanhua Du . Theoretical and experimental cocrystal screening of temozolomide with a series of phenolic acids, promising cocrystal coformers. Chinese Chemical Letters, 2024, 35(5): 109032-. doi: 10.1016/j.cclet.2023.109032
-
[7]
Beitong Zhu , Xiaorui Yang , Lirong Jiang , Tianhong Chen , Shuangfei Wang , Lintao Zeng . A portable and versatile fluorescent platform for high-throughput screening of toxic phosgene, diethyl chlorophosphate and volatile acyl chlorides. Chinese Chemical Letters, 2025, 36(1): 110222-. doi: 10.1016/j.cclet.2024.110222
-
[8]
Qingyun Hu , Wei Wang , Junyuan Lu , He Zhu , Qi Liu , Yang Ren , Hong Wang , Jian Hui . High-throughput screening of high energy density LiMn1-xFexPO4 via active learning. Chinese Chemical Letters, 2025, 36(2): 110344-. doi: 10.1016/j.cclet.2024.110344
-
[9]
Yao Zou , Difei Gong , Haiguang Yang , Hongmei Yu , Guorong He , Ningbo Gong , Lianhua Fang , Guanhua Du , Yang Lu . Prediction, screening, characterization, antioxidant and antihypoxic effects of multi-component zwitterionic cocrystals of dietary flavonoids with picolinic acid. Chinese Chemical Letters, 2025, 36(9): 110768-. doi: 10.1016/j.cclet.2024.110768
-
[10]
Yue Cao , Yue Lin , Li Chen , Huimin Niu , Renli Wei , Shuqian Qiu , Anjie Wang , Xiaoai Cao , Xiaofeng Lai , Yongshou Chen , Juanjuan Lin , Shuiliang Wang , Zhenyu Lin , Shenghang Zhang . Screening of glial fibrillary acidic protein specific aptamer and application in the development of fluorescent biosensor based on isothermal amplification strategy. Chinese Chemical Letters, 2026, 37(5): 111616-. doi: 10.1016/j.cclet.2025.111616
-
[11]
Hualei Xu , Manman Han , Haiqiang Liu , Liang Qin , Lulu Chen , Hao Hu , Ran Wu , Chenyu Yang , Hua Guo , Jinrong Li , Jinxiang Fu , Qichen Hao , Yijun Zhou , Jinchao Feng , Xiaodong Wang . 4-Nitrocatechol as a novel matrix for low-molecular-weight compounds in situ detection and imaging in biological tissues by MALDI-MSI. Chinese Chemical Letters, 2024, 35(6): 109095-. doi: 10.1016/j.cclet.2023.109095
-
[12]
Zhenxing Li , Yue Ding , Xinxin Tuo , Jinhong Hu , Taihong Zhang , Xiang Zhou , Liwei Liu , Song Yang . Structure-based drug repurposing targeting pathogenic virus superfamily 1 helicase: An integrated multi-computational screening and bioactivity identification strategy. Chinese Chemical Letters, 2025, 36(9): 110737-. doi: 10.1016/j.cclet.2024.110737
-
[13]
Wenzheng Li , Ke Zhang , Wenjing Liu , Ting Li , Wei Li , Lijuan Wu , Maodong Wang , Hangyun He , Pengfei Tu , Yuelin Song . Multidimensional host-guest molecule screening via 96-well plate assays to facilitate the development of natural product nanomicelles. Chinese Chemical Letters, 2026, 37(3): 111548-. doi: 10.1016/j.cclet.2025.111548
-
[14]
Yahui Li , Quanchen Feng , Krisztina László , Ying Wang . High-throughput screening of high C/N-ratio homonuclear dual-atom catalysts for electrochemical reduction of nitrate to ammonia. Chinese Chemical Letters, 2026, 37(3): 111536-. doi: 10.1016/j.cclet.2025.111536
-
[15]
Yanyan Yuan , Niu Zhang , Pangkuan Chen . An unprecedented supramolecular docking strategy enables rapid structure-determination of long alkyl-chain compounds. Chinese Journal of Structural Chemistry, 2025, 44(12): 100721-100721. doi: 10.1016/j.cjsc.2025.100721
-
[16]
Yaojun Li , Yun Li , Shenglong Liao , Yang Li , Shouchun Yin . Revolutionizing cancer therapies with organic photovoltaic non-fullerene acceptors: A deep dive into molecular engineering for advanced phototheranostics. Chinese Chemical Letters, 2025, 36(8): 110832-. doi: 10.1016/j.cclet.2025.110832
-
[17]
Jianning Zhang , Yihuai Zhang , Guoxin Ma , Jingchen Zhao , Tao Zhang , Jian Liu . Enhancing hydrothermal stability in Cu/SSZ-13 catalyst for diesel SCR applications through a novel core-shell structure. Chinese Chemical Letters, 2025, 36(7): 110516-. doi: 10.1016/j.cclet.2024.110516
-
[18]
Wenwen Zhang , Peichao Zhang , Conghao Gai , Xiaoyun Chai , Yan Zou , Qingjie Zhao . Unveiling Kinetics at Natural Abundance: 13C NMR Isotope Effect Experiments. University Chemistry, 2025, 40(10): 203-207. doi: 10.12461/PKU.DXHX202411076
-
[19]
Chen Chen , Jinzhou Zheng , Chaoqin Chu , Qinkun Xiao , Chaozheng He , Xi Fu . An effective method for generating crystal structures based on the variational autoencoder and the diffusion model. Chinese Chemical Letters, 2025, 36(4): 109739-. doi: 10.1016/j.cclet.2024.109739
-
[20]
Xiaoli Zhong , Liangsheng Chen , Hao Xu , Tianhang Jiang , Zhengyi Hua , Fancheng Tan , Xiaoya Mao , Ziquan Fan , Zhiwei Li , Jun Zeng , Shu-Hai Lin . Development of a comprehensive computational pipeline for cardiolipin atlas in an intermittent fasting model. Chinese Chemical Letters, 2025, 36(12): 111027-. doi: 10.1016/j.cclet.2025.111027
-
[1]
Metrics
- PDF Downloads(2)
- Abstract views(1398)
- HTML views(37)

 Login In
Login In




 DownLoad:
DownLoad: